-
杨属(Populus L.)植物分布广泛,不仅具有较高的经济和生态价值,同时也是木本模式植物,在林木分子生物学研究等方面具有重要作用。然而,作为北方城市重要的道路绿化和城市景观树种,杨树在提高城市绿化水平的同时,也因其花粉在每年春季大量飘散而引起了一定的植源性污染问题。杨树雄株散落的花粉以空气为传播媒介,通过人体呼吸及皮肤接触造成呼吸道疾病和皮肤过敏等问题,危害着人体健康[1-3]。除更换树种外,目前关于花粉污染的应对策略主要还包括人工喷水增湿、增强地表对花粉的吸附能力等。另外,日益发展的生物技术推动了林木遗传育种进程,有望成为解决花粉等植源性污染的有效途径之一。目前关于杨树的研究主要聚焦于其材性及抗逆性等方面,而对杨树的生殖发育研究相对较少。
杨树童期较长且雌雄异株,不同于大多数由萼片、花瓣、心皮(雌蕊/雄蕊)、胚珠4轮结构组成的被子植物花,杨树的雌花和雄花均高度简化,其中,雄花着生于苞片腋间,由退化的盘状花被和雄蕊构成,花药为黄色或红色,一般情况下药隔无明显突出[4]。花药为四孢型,成熟的花药壁由一层表皮、一层内层、两层中层和单层绒毡层组成。表皮宿存,内层发育成纤维状增厚,中层是短命的,其中一层在小孢子阶段变为扁平,最后被挤碎,另一层则在花药开裂前解体;绒毡层细胞早期是单核的,而当花粉母细胞减数分裂时则成为双核或多核的,它们在花粉单核时期开始退化,至双核期消失[5]。杨树性别特异的花器官发育是一个复杂的过程,但与其他被子植物花发育类似,杨树的花发育也分为开花诱导、花的发端和花器官发育3个受众多基因调控的复杂过程,且它们具有相似的花器官特性基因[6]。
随着研究的不断深入,花器官形成的相关理论逐步完善,由最初的“ABC”模型假说发展成为“ABCDE”模型及四聚体模型,其中,A类和E类基因决定花萼特征,A、B和E类基因一起调控花瓣发育,B、C和E类基因共同决定雄蕊发育,C类和E类基因决定心皮发育,D类和E类决定胚珠发育[7-8]。杨树花发育的相关器官特异性基因的作用不全同于其他开花植物,近年来调控杨树花器官发育的多个ABCDE类基因功能被解析,如:A类基因中的AP1同源基因PTM1、PTM2和B类基因中的AP3同源基因,它们在杨树雌/雄株中的表达有所差异,可能参与了杨树的性别决定[9-10];B类基因中的PI的同源基因PdPI和C类基因PTAG1和PTAG2,与杨树花发育密切相关并且可能影响营养器官的形成和发育[6, 11-12];E类基因中的PTM3、PTM4及PTM6,在杨树雌/雄株中花发育过程中持续表达,在胚珠、花药原基中有着非常高的表达量[13]。
AGAMOUS (AG)是MADS-box基因中研究较为充分的一类基因,属于植物特有的MIKC型C类MADS-box基因,包括euAG/PLENA (PLE)谱系和AGAMOUS-like11(AGL11)谱系,可与其他基因互作共同控制雄蕊及心皮的发育[8, 14-17]。在拟南芥中,AG可控制拟南芥花发育后期的花粉形成[18-20]。另有研究表明,在拟南芥AG突变体的花器官四轮结构中出现了第三轮结构(雄蕊)转化为花瓣、第四轮结构(心皮)发育成一个新的花的情况,而第一、二轮的花结构(萼片及花瓣)与野生型拟南芥花器官并无差异[21-23]。另外,抑制AG会导致菠菜雄性不育[24]。而AGAMOUS (AG)及其同源基因在单性花发育的研究中鲜有报道,在毛果杨雌株中,AG(PtAG1及PtAG2)和STK(AGL11)基因在杨树种毛发育中表现出较强的功能保守性,共同抑制种毛的形成[12],而该基因在杨树雄花花发育中的作用尚无报道。
本研究基于AG2在被子植物花器官的决定作用,对银腺杨‘84K’(Populus alba × P. glandulosa ‘84K’)PagAG2基因时空特异性进行了检测,为进一步研究其在杨树雄花花发育中的功能,进而有针对性地对杨树生殖发育进行调控奠定基础。
-
银腺杨‘84K’(Populus alba × P. glandulosa ‘84K’)花序于2019年2月22日采自北京市中国林科院内(115.7°~117.4° E,39.4°~41.6° N),组培材料生长于温度25℃、光强2300 Lx、光周期16 h/8 h的组织培养间中;RNA提取试剂盒购自Aidlab公司(北京);Premix TaqTM酶购、反转录试剂盒、qRT-PCR试剂盒均购自Takara公司(日本);固定液由50%FAA固定液100 mL、无水乙醇50 mL、37%甲醛溶液10 mL、冰乙酸5 mL加DEPC·H2O定容至100 mL;RNA预杂交液/杂交液由20 × SSC 30 mL、50 × Denhardt's 10 mL、10%SDS 5 mL、10 mg· mL−1Salmon DNA 1 mL、Formamide(甲酰胺)50 mL、ddH2O 4 mL,充分混匀后使用0.45 μm滤膜去杂质后使用;抗荧光淬灭封片剂购自生工生物公司(上海)。
-
取‘84K’杨越冬花枝,2019年2月22日置于温室水培(0 d),每日更换一次水。对水培后1周内的花序连续采样(0、3、4、5、6、7 d),置于PBS固定液中立即真空抽气固定,并依次完成脱水、透明、浸蜡、包埋、修块、切片、粘片、脱蜡步骤,以番红—固绿法染色[25]。染色完成染色后,在光学显微镜(OLYMPUS公司(BX51)产品)下观察、拍照记录。
-
分别提取‘84K’杨组培苗的根、茎、叶及水培后2周内的花序连续采样(水培后0、3、4、5、6、7 d的花序)的总RNA,并反转录合成cDNA。使用荧光定量PCR仪(Roche公司(Light Cycle 480Ⅱ)产品)进行qRT-PCR反应,反应体系为cDNA模板2 μL,Primer-F(10 μmol·L−1) 0.8 μL,Primer-R(10 μmol · L−1) 0.8 μL,XTB Green Premix TaqII(TliRNaseH Plus,2 × )10 μL,ddH2O补足至20 μL。其扩增条件为预变性95℃,30 s;变性95℃,5 s,退火60℃,30 s,40个循环;熔解曲线为95℃ 5 s,65℃ 1 min。以Actin为内参基因,每个样品进行3次重复,用2−ΔΔCT算法进行分析[26],引物见表1。
-
基于碱基互补的原则,利用荧光原位杂交技术将细胞原位杂交技术和荧光技术有机结合,用荧光素标记的已知外源RNA作探针,与‘84K’杨雄花组织切片杂交,与待测核酸的靶序列专一性结合,通过检测杂交位点荧光来显示PtAG2核苷酸序列在‘84K’杨雄花中的位置。根据待检测目的基因序列,采用Primer Premier 6.0软件设计,并用探针合成仪器(UNICORN 5.31,沃特曼)合成RNA探针(表2),荧光素标记反义RNA核酸探针,以正义RNA探针为阴性对照。将‘84K’杨雄花序样品放入配制好的固定液(50%FAA固定液100 mL、无水乙醇50 mL、37%甲醛溶液10 mL、冰乙酸5 mL,加DEPC·H2O定容至100 mL)中固定12 h后经梯度酒精脱水后浸蜡、包埋、切片(SQ2125,徕克公司)、摊片(PPTHK-21B,徕克)、捞片,62℃烤箱烤片2 h。石蜡切片脱蜡后,将切片于修复液中煮沸10~15 min,自然冷却。滴加蛋白酶K(20 ug· mL−1)37℃消化,纯水冲洗后PBS洗3次,每次5 min。滴加预杂交液,37℃于杂交仪(S500-24,ThermoBrite)孵育1 h。倾去预杂交液,滴加探针杂交液,置于恒温箱57℃度杂交过夜。洗去杂交液,2×SSC,37℃洗10 min,1×SSC,37℃洗2次,每次5 min,0.5×SSC室温洗10 min。切片滴加DAPI染液,避光孵育8 min,冲洗后滴加抗荧光淬灭封片剂(Anti-Fade Mounting Medium,生工)封片,置于荧光显微镜下(CX41,OLYMPUS)观察并采集图像(紫外激发波长330~380 nm,发射波长420 nm,发蓝光;FAM(488)绿光激发波长465~495 nm,发射波长515~555 nm,发绿光;CY5红光激发波长510~560 nm,发射波长590 nm,发红光)。所有试剂、仪器均需DEPC处理。
表 1 qRT-PCR表达分析引物
Table 1. Primers used in qRT-PCR analysis
基因 Gene 产物长度 Amplification Length/bp 引物序列 Primer sequence 用途 Usage Actin 195 F: 5′-AAACTGTAATGGTCCTCCCTCCG -3′ Internal
referenceR: 5′-GCATCATCACAATCACTCTCCGA -3′ PagAG2 270 F: 5′-GCTTCTCCGAGCAAAGGTCT-3′ qRT-PCR R: 5′-GCAGGCGGCAATCCATTAAC -3′ 表 2 RNA探针序列
Table 2. RNA probe sequences
基因 Gene 序列 Sequence 荧光标记 Fluorescence labeling PagAG2 5'-AUUAUCUGUUCUUUGCGAUGCUGAGGUUGC-3' 5’CY5修饰 红色
(阴性对照)5'-UAAUAGACAAGAAACGCUACGACUCCAACG-3' 5’FAM修饰 绿色 -
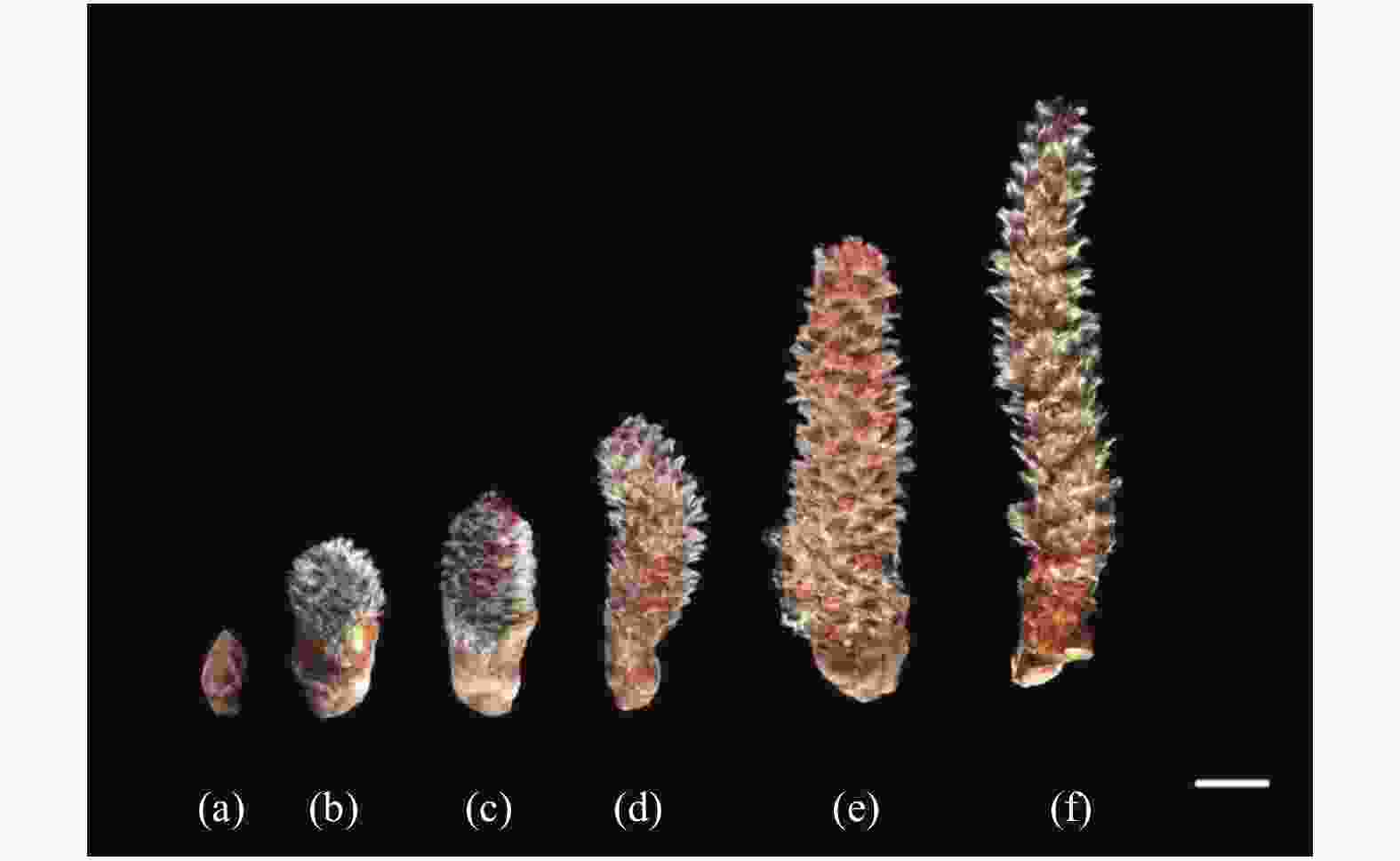
杨树雄花为柔荑花序,着生于苞片腋间,雄蕊被退化的盘状花被所包围。‘84K’杨越冬花枝自水培之日(0 d)起,其雄花花芽迅速膨大并将苞片撑开,整个花序覆有白色绒毛(3 d),第4天可见红色花药,之后整个花序继续伸长,随着花药的成熟花药颜色由红转为黄,由于花发育的顺序由下向上,花序轴上端的花和下部的花的发育时期相差较大,雄花开始有部分花粉散出,至7 d时,花粉大量散出,苞片脱落。水培后第0、3、4、5、6、7天的花序度(3个样品)测量均值分别为0.8、2.5、3.1、3.7、6.5、8 cm(图1)。
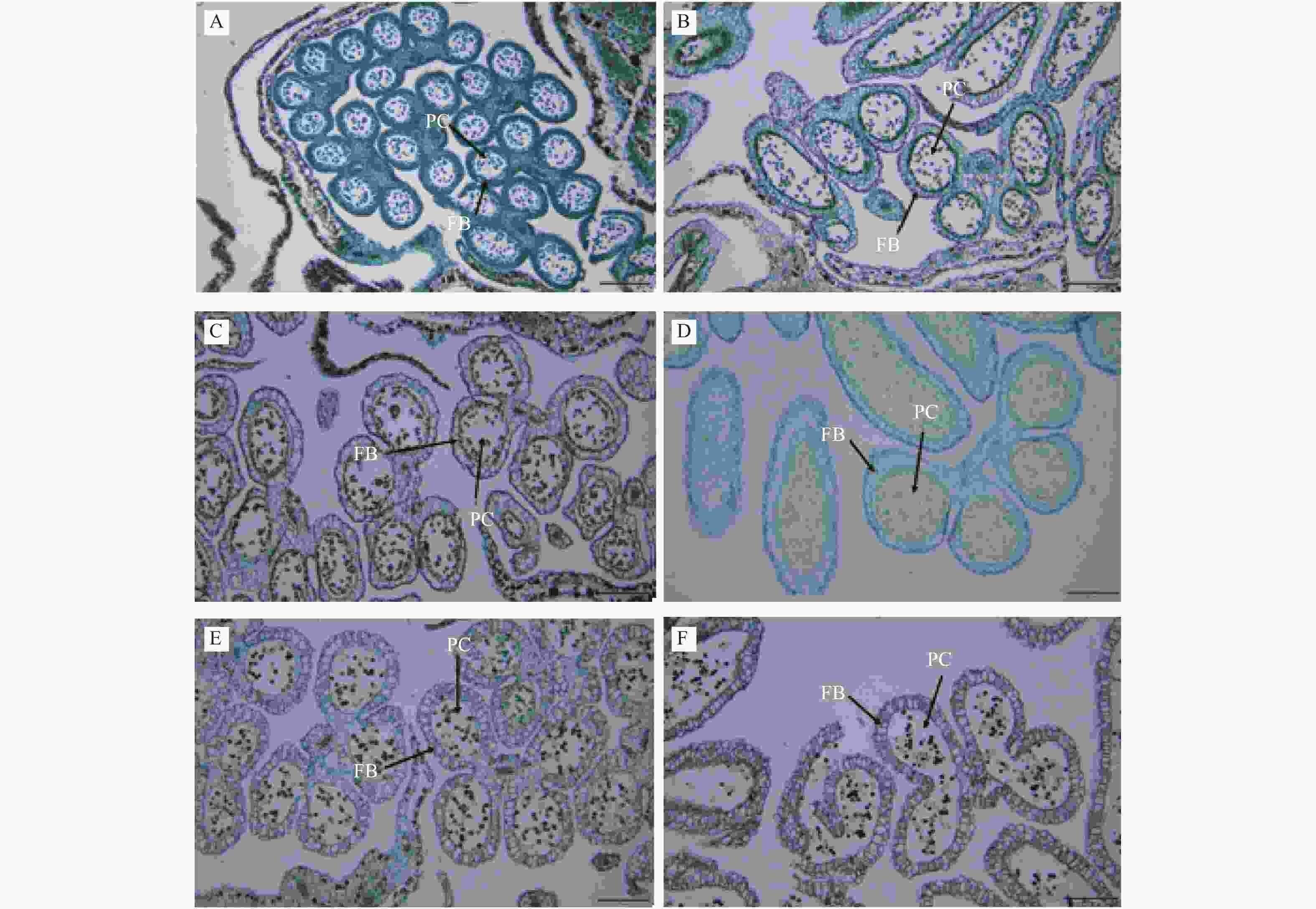
从石蜡切片结果看出:取样时杨树雄花花药中的单倍体的小孢子已从四分体中释放,游离于花粉囊中,取样0~7 d随花药横切面增大,纤维状带逐渐展开,绒毡层浓缩继而解体,中层降解,至7 d时药隔每侧的2个花粉囊互相连通,待花粉囊开裂出现花药裂口,成熟的花粉依而从花粉囊内散出而传粉(图2)。
-
利用qRT-PCR技术检测了PagAG2在‘84K’杨不同器官及其雄花花序不同发育时期的表达差异。结果表明,PagAG2在雄花发育的过程中,水培后1周内的花序样品中PagAG2的表达量逐渐升高后降低,6 d达到顶峰(图3A),之后回落,7 d可见PagAG2降至整个观察期最低,药隔两侧花粉囊互相连通、花粉成熟将大量散开花粉,这一过程十分迅速,PagAG2基因与杨树雄花花发育密切相关。然而,值得注意的是,PagAG2在根、茎、叶这些营养器官中也有一定表达(图3B),并且在茎中的表达量高于根和叶,但均低于整个观察期的雄花。
-
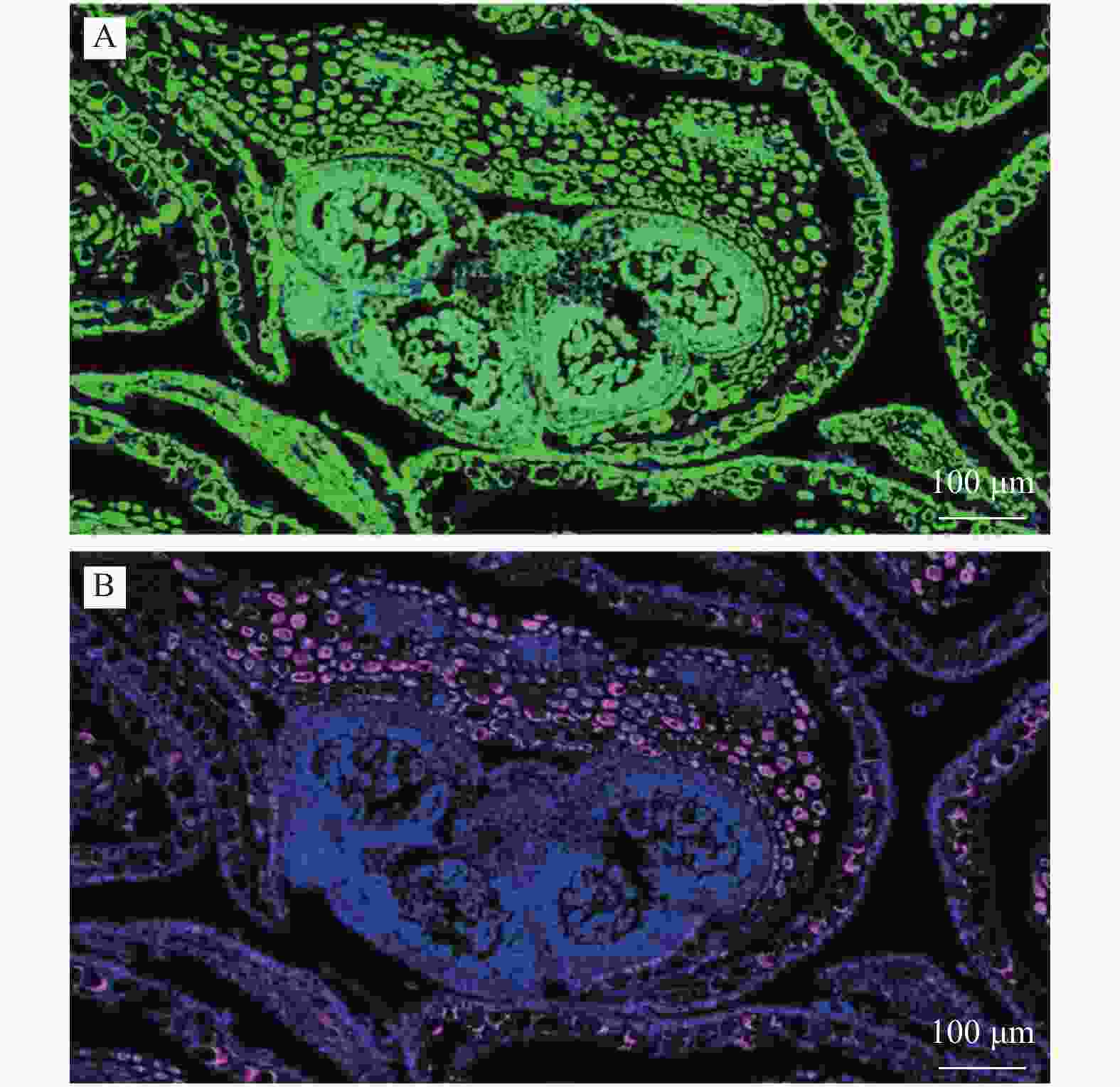
通过原位杂交的技术手段检测了PagAG2在杨树雄花中的具体表达部位。用PagAG2特异性探针对‘84K’杨雄花花序进行原位杂交,洗脱后,其荧光显微镜镜检拍照结果见图4。在样本中可以清晰地观察到亮绿色杂交信号覆盖整个视野中的花序组织,而红色阴性对照仅出现在花药周围的花被组织。PagAG2在‘84K’杨雄花花药中特异性表达。
-
杨树花发育过程涉及大量的基因调控,C类基因AG编码的MADS-box转录因子在其中起着关键作用,包括终止花分生组织的生成、调控雄蕊/心皮/胚珠的发育,并与A类基因相互拮抗[27-31]。Brunner等[11]的研究表明,PtAG2在毛白杨雌株和雄株中的表达模式一致,表明它们不太可能在功能上与树的性别相关,但却对杨树雌/雄花相关组织的形成和发育有着重要作用。本试验中,PagAG2在‘84K’杨雄花花序中的表达具有时空特异性。通过实时定量PCR检测水培后0~7 d的‘84K’杨雄花花序PagAG2基因表达量,时间上PagAG2在花组织中的相对表达量先增加后降低,从对应的雄花花序外部形态及石蜡切片上看,其表达量是随着雄花花序发育而逐渐升高,在花药成熟时达到顶峰,并在花粉成熟散落后降低。另外,本试验通过实时定量检测了PagAG2‘84K’杨根、茎、叶的表达量,利用荧光原位杂交展现了PagAG2花序中的表达部位,空间上PagAG2的转录水平在杨树花序表达量高于营养器官,并且花粉为PagAG2在花序中具体表达部位,在花序中的整个花药均有表达。PagAG2表达与‘84K’杨雄花发育密切相关,有望作为基因工程改良杨树花粉的目标基因。植物生殖发育的整个过程在一定程度上均受到MADS-box基因调控[32]。另外,已有研究表明,MADS-box蛋白在杨树营养器官生长发育中也发挥重要作用,如杨树SVL参与营养芽休眠过程,PTM5、ADF、LRR、FPF1被证明能够调控木质部的形成等[9, 33-37]。目的基因PagAG2除可能参与杨树雄花花发育,还可能参与杨树营养组织的生长发育调控。实时定量PCR试验中,‘84K’杨幼期根、茎、叶营养组织中也检测到了PaAG2转录物,但其转录水平低于试验材料自水培日起到花粉成熟并大量散落的整个观察期的花组织。已有研究表明,AG表达的水平对于确定器官身份是重要的[38],AG的表达受其它基因的调节,如CURLY LEAF能够阻止营养期AG在叶中表达[39],Brunner等[11]推测,PtAG2营养表达可能是杨树中不太严格的抑制控制的结果,但该基因是否参与杨树其他组织的某些生长发育调控,需要对其做进一步的基因功能分析。
杨树基因组几乎包含了拟南芥所有调控开花的主要同源基因[40-41],在对模式植物拟南芥的研究中,发现了促进花器官发育的多聚MADS-box蛋白复合物的形成。MADS-box蛋白以二聚体的形式与下游靶基因结合,实现对拟南芥生长发育的调控作用,如AG、AP2和AGL1在拟南芥中能够形成同源二聚体蛋白共同决定花器官特性[42-43]。随着研究的深入又提出了四分子模型,即2个MADS蛋白形成二聚体,又分别与靶基因结合后形成蛋白四聚体,共同对花器官的建成起作用,四聚体PI-AP3-SEP3-AG控制雄蕊的分化,过表达PI-AP3-SEP3-AG植物的营养叶转化为雄蕊状器官[44],这也为其他物种中MADS-box基因相关调控机理的研究提供了参考,杨树PagAG2也可能与其他它因形成多聚体,共同实现复杂的调控作用。
-
本研究对‘84K’杨PagAG2基因的表达模式进行了研究。时间上PagAG2表达量随着花粉成熟先增加后降低,空间上PagAG2在杨树花序表达量高于营养器官,并且花粉为PagAG2在花序中具体表达部位,PagAG2与‘84K’杨雄花花发育密切相关,有望作为基因工程改良杨树花粉的目标基因,而PagAG2的转录物在杨树幼期根、茎、叶组织中也能被检测得到,其是否参与杨树其他组织的生长发育调控,在相关育种工作中需要对该基因做进一步的基因功能分析,以便利用基因编辑等技术沉默杨树花发育相关基因PagAG2,产生对营养生长无副作用的少花粉、无花粉表型,从而为从源头上解决杨树花粉过敏的问题提供一种有效途径。
银腺杨PagAG2基因的时空表达分析
Spatiotemporal Expression Analysis of PagAG2 Gene in Populus alba × P. glandulosa '84K'
-
摘要:
目的 研究杨树AG类基因(PtAG1及PtAG2)在杨树雄花发育过程中的作用,为杨树分子育种提供基础。 方法 以‘84K’杨(Populus alba × P. glandulosa ‘84K’)花芽为材料,对花芽发育过程连续采样,观察表型发育状况与测定PagAG2基因部位与表达量,通过石蜡切片、实时定量PCR、原位杂交等技术手段,对银腺杨‘84K’(Populus alba × P. glandulosa ‘84K’)PagAG2基因时空特异性进行研究,分析PagAG2基因表达模式。 结果 在‘84K’杨雄花发育过程中,花药逐渐成熟,在此过程中,PagAG2的表达量先增加后降低,并且在‘84K’杨组培苗根、茎、叶中也能够检测到该转录物。原位杂交结果显示:PagAG2在‘84K’杨雄花花药中表达,但在包围花药的周围组织中不表达。 结论 ‘84K’PagAG2基因与‘84K’杨雄花花发育密切相关,有望作为基因工程改良杨树花粉的目标基因用于研究。但PagAG2是否参与杨树其他组织的生长发育调控,在相关育种工作中需要对其做进一步的基因功能分析。 Abstract:Objective To identify the role of PtAG2 gene in the development of male flowers of poplar. Method Based on the regulation of AG genes (PtAG1 and PtAG2) in the stamens and carpels of monoecious plants, the spatial and temporal specificity of PagAG2 gene of Populus alba × P. glandulosa '84K' was studied by means of paraffin section, real-time quantitative PCR and in situ hybridization. Taking the flower branches of P. alba × P. glandulosa '84K' as test materials, the inflorescences were continuously sampled within one week after hydroponic culture to observe the development of male flowers by paraffin sectioned technique. Real-time quantitative PCR was used to detect the expression of PagAG2 in the roots, stems, leaves and inflorescence of P. alba × P. glandulosa '84K'. In situ hybridization was used to detect the specific expression sites of PagAG2 in the flower organs of P. alba × P. glandulosa '84K' male flowers, so the expression pattern of PagAG2 gene was analyzed. Result During the development of male flowers of P. alba × P. glandulosa '84K', the anthers matured gradually. At the same time, the expression of PagAG2 increased at initial and then decreased, and the transcription was also be detected in the roots, stems and leaves of tissue cultured P. alba × P. glandulosa '84K'. The results of in situ hybridization showed that PagAG2 was expressed in the anthers of P. alba × P. glandulosa '84K' male flowers but not in the surrounding tissues. Conclusion PagAG2 gene is closely related to the male flower development of P. alba × P. glandulosa '84K', so it is expected to be a target gene for genetic engineering to improve poplar pollen. However, whether the PagAG2 gene participates in the regulation of the growth and development of other tissues of poplar needs to be further studied. -
Key words:
- Populus alba × P. glandulosa '84K'
- / pollen
- / PagAG2
-
表 1 qRT-PCR表达分析引物
Table 1. Primers used in qRT-PCR analysis
基因 Gene 产物长度 Amplification Length/bp 引物序列 Primer sequence 用途 Usage Actin 195 F: 5′-AAACTGTAATGGTCCTCCCTCCG -3′ Internal
referenceR: 5′-GCATCATCACAATCACTCTCCGA -3′ PagAG2 270 F: 5′-GCTTCTCCGAGCAAAGGTCT-3′ qRT-PCR R: 5′-GCAGGCGGCAATCCATTAAC -3′ 表 2 RNA探针序列
Table 2. RNA probe sequences
基因 Gene 序列 Sequence 荧光标记 Fluorescence labeling PagAG2 5'-AUUAUCUGUUCUUUGCGAUGCUGAGGUUGC-3' 5’CY5修饰 红色
(阴性对照)5'-UAAUAGACAAGAAACGCUACGACUCCAACG-3' 5’FAM修饰 绿色 -
[1] 杨 颖, 王 成, 李 伟. 空气花粉污染研究的现状与趋势[J]. 中国城市林业, 2007, 5(2):52-53. doi: 10.3969/j.issn.1672-4925.2007.02.020 [2] 杨 颖, 王 成, 郄光发. 城市植源性污染及其对人的影响[J]. 林业科学, 2008, 44(4):151-155. doi: 10.3321/j.issn:1001-7488.2008.04.027 [3] 李 倩, 靳 颖, 华振玲, 等. 空气致敏花粉污染研究进展[J]. 生态学报, 2005, 25(2):334-338. doi: 10.3321/j.issn:1000-0933.2005.02.023 [4] 洪 涛, 麻左力, 陈敬诗. 大叶杨花部形态及其在杨属的分类位置[J]. 植物学报, 1987, 29(3):236-241. [5] 李文钿, 马丰山. 木本植物有性杂交生殖生物学图谱[M]. 北京: 科学出版社, 2011: 15-16. [6] 张冰玉, 苏晓华, 周祥明. 杨树花发育相关基因及基因工程调控[J]. 分子植物育种, 2007,5(5):695-700. doi: 10.3969/j.issn.1672-416X.2007.05.016 [7] Coen E S, Meyerowitz E M. The war of the whorls: genetic interactions controlling flower development[J]. Nature, 1991, 353(6339): 31-37. doi: 10.1038/353031a0 [8] Theien G. Development of floral organ identity: Stories from the MADS house[J]. Current Opinion in Plant Biology, 2001, 4(1): 75-85. doi: 10.1016/S1369-5266(00)00139-4 [9] Cseke L J, Zheng J, Podila G K. Characterization of PTM5 in aspen trees: a MADS-box gene expressed during woody vascular development[J]. Gene, 2003, 318(1): 55-67. [10] Sheppard L A, Brunner A M, et al. A DEFICIENS homolog from the dioecious tree black cottonwood is expressed in female and male floral meristems of the two-whorled, unisexual flowers[J]. Plant Physiol, 2000, 124(2): 627-640. doi: 10.1104/pp.124.2.627 [11] Brunner A M, Rottmann W H, Sheppard L A, et al. Structure and expression of duplicate AGAMOUS orthologues in poplar[J]. Plant Molecular Biology, 2000, 44(5): 619-634. doi: 10.1023/A:1026550205851 [12] Lu H, Klocko A L, Brunner A M, et al. RNAi suppression of AGAMOUS and SEEDSTICK alters floral organ identity and impairs floral organ determinacy, ovule differentiation, and seed-hair development in Populus[J]. New Phytologist, 2019, 222(2): 923-937. doi: 10.1111/nph.15648 [13] Cseke L J, Cseke S B, Ravinder N, et al. SEP-class genes in Populus tremuloides and their likely role in reproductive survival of poplar trees[J]. Gene, 2005, 358(1): 1-16. [14] Kramer E M. Patterns of gene duplication and functional evolution during the diversification of the AGAMOUS subfamily of MADS box genes in angiosperms[J]. Genetics, 2004, 166(2): 1011-1023. doi: 10.1093/genetics/166.2.1011 [15] Zahn L M, Leebens-Mack J H, Arrington J M, et al. Conservation and divergence in the AGAMOUS subfamily of MADS‐box genes: evidence of independent sub‐ and neofunctionalization events[J]. Evolution & Development, 2006, 8(1): 30-45. [16] Dreni L, Kater M M. MADS reloaded: evolution of the AGAMOUS subfamily genes[J]. New Phytologist, 2014, 201(3): 717-732. doi: 10.1111/nph.12555 [17] Honma T, Goto K. Complexes of MADS-box proteins are sufficient to convert leaves into floral organs[J]. Nature, 2001, 409(6819): 525-529. doi: 10.1038/35054083 [18] Ito T, Wellmer F, Yu H, et al. The homeotic protein AGAMOUS controls microsporogenesis by regulation of SPOROCYTELESS[J]. Nature, 2004, 430(6997): 356-60. doi: 10.1038/nature02733 [19] Ito T, Ng K H, Lim T S, et al. The homeotic protein AGAMOUS controls late stamen development by regulating a jasmonate biosynthetic gene in Arabidopsis[J]. Plant Cell, 2007, 19(11): 3516-3529. doi: 10.1105/tpc.107.055467 [20] Wilson Z A, Zhang D B. From Arabidopsis to rice: pathways in pollen development[J]. J Exp Bot, 2009, 60(5): 1479-1492. doi: 10.1093/jxb/erp095 [21] Bowman J L. Genes directing flower development in Arabidopsis[J]. Plant Cell, 1989, 1(1): 37-52. [22] Yanofsky M F, Ma H, Bowman J L, et al. The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors[J]. Nature, 1990, 346(6279): 35-39. doi: 10.1038/346035a0 [23] Dennis L, Peacock J. Genes directing flower development in Arabidopsis[J]. Plant Cell, 2019, 31(6): 1192-1193. doi: 10.1105/tpc.19.00276 [24] Sather D N, Jovanovic M, Golenberg E M. Functional analysis of B and C class floral organ genes in spinach demonstrates their role in sexual dimorphism[J]. BMC Plant Biol, 2010, 10: 46. doi: 10.1186/1471-2229-10-46 [25] 李正理. 植物组织制片学[M]. 北京: 北京大学出版社, 1996: 130-138. [26] Sun J, Niu Q W, Tarkowski P, et al. The ArabidopsisAtIPT8/PGA22 gene encodes an isopentenyl transferase that is involved in De Novo cytokinin biosynthesis[J]. Plant Physiology, 2003, 131(1): 167-176. doi: 10.1104/pp.011494 [27] 高志红, 张玉明, 王 珊, 等. 植物花发育调控基因AGAMOUS 的研究进展[J]. 西北植物学报, 2008, 28(3):638-644. doi: 10.3321/j.issn:1000-4025.2008.03.034 [28] 龚霞峰, 胡江琴, 刘姬艳, 等. 植物AGAMOUS同源基因的表达调控[J]. 杭州师范大学学报:自然科学版, 2009, 8(3):218-223. [29] 王 雪, 田京京, 李 爽, 等. AGAMOUS在花发育中的核心功能研究进展[J]. 分子植物育种, 2020, 18(9):2878-2885. [30] Angenent G C, Franken J, Busscher M, et al. A novel class of MADS box genes is involved in ovule development in petunia[J]. The Plant Cell, 1995, 7(10): 1569-1582. [31] Cronk Q, Soolanayakanahally R, Bräutigam K. Gene expression trajectories during male and female reproductive development in balsam poplar (Populus balsamifera L.)[J]. Sci Rep, 2020, 10(1): 8413. doi: 10.1038/s41598-020-64938-w [32] Kaufmann K, Melzer R, Theissen G. MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants[J]. Gene, 2005, 347(2): 183-198. doi: 10.1016/j.gene.2004.12.014 [33] Shim D, Ko J H, Kim W C, et al. A molecular framework for seasonal growth-dormancy regulation in perennial plants[J]. Hortic Res, 2014, 1(1): 14059. doi: 10.1038/hortres.2014.59 [34] Busov V B. Plant development: dual roles of Poplar SVL in vegetative bud dormancy[J]. Curr Biol, 2019, 29(2): R68-R70. doi: 10.1016/j.cub.2018.11.061 [35] Cseke L J, Ravinder N, Pandey A K, et al. Identification of PTM5 protein interaction partners, a MADS-box gene involved in aspen tree vegetative development[J]. Gene, 2007, 391(1-2): 209-222. doi: 10.1016/j.gene.2006.12.033 [36] Hoenicka H, Lautner S, Klingberg A, et al. Influence of over-expression of the Flowering Promoting Factor 1 gene (FPF1) from Arabidopsis on wood formation in hybrid poplar (Populus tremula L. × P. tremuloides Michx.)[J]. Planta, 2012, 235(2): 359-373. doi: 10.1007/s00425-011-1507-8 [37] Leseberg CH, Li A, Kang H, et al. Genome-wide analysis of the MADS-box gene family in Populus trichocarpa[J]. Gene, 2006, 378(1): 84-94. [38] Mizukami Y, Ma H. Separation of AG function in floral meristem determinacy from that in reproductive organ identity by expressing antisense AG RNA[J]. 1995, 28(5): 767-784. [39] Goodrich J, Puangsomlee P, Martin M, et al. A Polycomb-group gene regulates homeotic gene expression in Arabidopsis[J]. Nature, 1997, 386(6620): 44-51. doi: 10.1038/386044a0 [40] Chen Z, Rao P, Yang X, et al. A global view of transcriptome dynamics during male floral bud development in Populus tomentosa[J]. Sci Rep, 2018, 8(1): 722. doi: 10.1038/s41598-017-18084-5 [41] Brunner A M, Nilsson O. Revisiting tree maturation and floral initiation in the poplar functional genomics era[J]. New Phytologist, 2010, 164(1): 43-51. [42] Riechmann J L, Meyerowitz E M. Determination of floral organ identity by Arabidopsis MADS domain homeotic proteins AP1, AP3, PI, and AG is independent of their DNA-binding specificity[J]. Mol Biol Cell, 1997, 8(7): 1243-1259. doi: 10.1091/mbc.8.7.1243 [43] Fan H Y, Hu Y, Tudor M, et al. Specific interactions between the K domains of AG and AGLs, members of the MADS domain family of DNA binding proteins[J]. Plant J, 1997, 12(5): 999-1010. doi: 10.1046/j.1365-313X.1997.12050999.x [44] Favaro R, Pinyopich A, Battaglia R, et al. MADS-box protein complexes control carpel and ovule development in Arabidopsis[J]. Plant Cell, 2003, 15(11): 2603-2611. doi: 10.1105/tpc.015123 -



 下载:
下载: