-
质体蓝素(Phytocyanin,PC)是一类存在于植物中的古老超蛋白家族,该家族均有质体蓝素样结构域(Plastocyanin-like domain,PCLD),具有2个能形成二硫键的半胱氨酸(Cys),其中, 部分成员含有4个保守的铜离子结合位点:2个组氨酸(His)、1个半胱氨酸和1个甲硫氨酸(Met)或谷氨酰胺(Gln)[1-2]。根据铜离子结合位点的氨基酸残基类型、光谱和氧化还原性能、蛋白结构组分等特征,PCs家族分为4个亚家族:花青苷样蛋白(Uclacyanin-like protein,UCL)、漆树兰蛋白(Stellacyanin-like protein,SCL)、质体蓝素样蛋白(Plantacyanin-like protein,PLCL)和早期结瘤素样蛋白(Early nodulin-like protein,ENODL)[3]。UCL和PLCL有相同的铜离子结合位点(2个His,1个Cys和1个Met),但UCL属于嵌合型糖蛋白,其蛋白骨架中含有糖蛋白样结构域,而PLCL为非糖蛋白[3]。SCL的铜离子结合位点中Gln替代了Met,SCL和UCL家族不仅包含天冬酰胺(Asn)残基连接的N-糖基化位点, 还具有通过丝氨酸(Ser)和羟脯氨酸(Hyp)残基连接的O-糖基化位点[4]。ENODL与其它亚家族之间有高度的相似性,但该亚家族成员的蛋白骨架缺完整的铜离子结合位点,铜配位体中的4个氨基酸残基部分或全部被其它氨基酸残基取代[5]。研究发现,拟南芥(Arabidopsis thaliana)和水稻(Oryza sativa)中的大多数ENODLs是嵌合的阿拉伯半乳聚糖蛋白,其蛋白骨架包含N-端信号肽(Signal peptide,SP)和阿拉伯半乳聚糖蛋白样域(Arabinogalactan protein-like region,ALR)[5-6]。
PC家族中能结合铜离子的3个亚家族(UCL、SCL和PLCL)为电子传递体。研究显示,PCs在植物发育和抗逆过程扮演着重要角色[5, 7-8],PCs参与了植物对非生物胁迫的应激反应,如干旱胁迫、盐胁迫、冷胁迫和金属离子等[5, 9-11]。
全基因组分析是鉴定基因家族成员、阐明其生物学作用的首要步骤和有效途径。利用生物信息学分析方法对植物PC家族基因全基因组分析的报道不多,仅在拟南芥、水稻和大白菜(Brassica rapa)基因组中分别发现38、62和84条PCs基因[5-7]。铁皮石斛(Dendrobium officinale)为兰科石斛属植物中具有观赏价值和医药价值的一种附生兰,其基因组测序的完成为该物种PC基因家族(DoPC)的全基因组预测和生物信息学分析提供了宝贵资源,有助于了解该基因家族的潜在功能。本研究可为石斛属PC基因家族功能验证提供数据基础。
-
铁皮石斛基因组数据库:http://202.203.187.112/herbalplant;拟南芥基因组数据库:http://arabidopsis.org/;国家水稻数据中心数据库:http://www.ricedata.cn/gene/;SMART数据库:http://smart.embl-heidelberg.de;pfam数据库:http://pfam.xfam.org/。
-
氨基酸序列分析软件:BLASTP,clustalW;N-端信号肽预测:SignalP 4.1(http://www.cbs.dtu.dk/services/SignalP/);糖基磷脂酰肌醇锚定信号(C-terminal glycosylphosphatidylinositol signal,GAS)预测:Big-PI Plant Predictor(http://mendel.imp.ac.at/gpi/plant_server.html)和PSORT(http://psort.hgc.jp/form.html);N-糖基化位点预测:NetNGlyc 1.0 Server(http://www.cbs.dtu.dk/services/NetNGlyc/);系统进化分析软件:MEGA 5.0;图像处理软件:PhotoShop。
-
下载铁皮石斛全基因组蛋白编码序列和拟南芥、水稻PC基因家族成员的蛋白序列(AtPCs和OsPCs)到本地。利用BLASTP工具进行比对搜索(缺省参数设置),手动去除错配序列。
-
将BLASTP比对获得的潜在DoPCs蛋白序列文件保存为FASTA格式,利用SMART和pfam数据库验证DoPCs蛋白序列的质体蓝素样结构域PCLD(缺省参数设置)。
-
将存在PCLD结构域的DoPCs蛋白序列文件保存为FASTA格式,利用SignalP 4.1、Big-PI Plant Predictor和NetNGlyc 1.0 Server软件分别预测其SP,GAS位点、N-糖基化位点(缺省参数设置)。依据Mashifuchi等[6]研究拟南芥AtPCs蛋白序列潜在ALR的原则:以非邻接的脯氨酸(Pro)残基基序([Ala/Ser/Thr/Gly]-Pro-X(0, 10)-[Ala/Ser/Thr/Gly]-Pro)和邻接的Pro残基基序([Ala/Ser/Thr/Gly]-Pro3-4)为阿拉伯半乳聚糖(AG)糖模块;Ser-Pro2-4为假定的伸展蛋白糖模块,手动分析DoPCs蛋白序列中潜在的ALRs。
-
利用软件Clustal W 2.1对DoPCs氨基酸序列PCLD结构域进行多重序列比对分析(缺省参数设置),手动标记结构域特征位点。
-
首先,利用Clustal W软件对所有DoPCs的氨基酸序列进行比对,生成软件MEGA可读的比对文件(缺省参数设置);然后,利用MEGA5.0软件采用邻接法(Neighbor-joining Method)对上述文件进行进化树分析。Bootstrap设置为1 000。最后生成无根树。
-
按照Zhao等[12]发表的方法采集正常生长的铁皮石斛成熟种子(中国林业科学研究院科研温室),分为2组进行体外共生和非共生萌发试验。一组种子用来构建与美孢胶膜菌(Tulasnella calospora, ITS序列GenBank登录号:GU166418.1)体外共生萌发体系(DoTcg),选取萌发到第三阶段(原分生组织出现)的种子利用70%酒精表面消毒后进行台盼蓝染色[13],在种子内检测到共生真菌菌丝,表明共生体系建立[13];另一组种子进行体外非共生萌发到相同阶段(Do),两组试验分别设置三组生物学重复。液氮取样、提取总RNA、构建RNA-seq文库进行高通量测序[12]。将测序获得的DoPCs家族基因reads数进行标准化处理获得基因表达的RPKM值(Reads per kilobase per million mapped reads)值[12],再取log2对数输入软件HemI v1.0(Heatmap Illustrator)绘制DoPCs基因表达热图[14]。设置|log2(Fold-change)| > 1(p-value < 0.01)作为基因差异表达筛选条件。
-
利用BLASTP搜索比对工具从铁皮石斛蛋白数据库中分别找出与拟南芥PCs(AtPCs)和水稻(OsPCs)相似度较高的序列(E-value < 0.001),经过SMART预测和Pfam数据库比对两种方法验证,在拟南芥和水稻中均有38条PC样蛋白序列(DoPCs)含有1个PCLD结构域。已知拟南芥基因组中含有38条PC基因[6, 15],在数量上与铁皮石斛的PC基因相同;而水稻和大白菜基因组中分别含有62条和84条PC基因,远多于铁皮石斛和拟南芥基因组中PC基因的数量。
对DoPCs家族成员的PCLD结构域进行多重序列比对分析,结果(图 1)表明:38条DoPCs蛋白序列均含有2个Cys残基,Cys在DoPCs家族中高度保守;19条DoPCs蛋白序列中含有4个完整的铜结合配体(His、Cys、His、Met/Gln),其中,11条为His、Cys、His、Met类型,8条为His、Cys、His、Gln类型。19个可结合铜的DoPCs蛋白归类为3个亚家族中:5个花青苷样蛋白(DoUCLs)、8个漆树蓝样蛋白(DoSCLs)和6个质体蓝素样蛋白(DoPLCLs);其余19个不含铜结合位点的DoPCs为早期结瘤素样蛋白(DoENODLs)(表 1)。多重序列比对发现在DoENODLs亚家族中,与其它亚家族的铜结合位点所对应的位置上有一些较保守的基序,这些氨基酸残基取代了His、Cys、His、Met/Gln,可能与结合铜有关。用亚家族名称命名后的DoPCs所对应的铁皮石斛基因ID、结构域数量及铜结合位点等配体信息见表 1。
表 1 DoPC家族成员及其结构信息
Table 1. The putative DoPCs named information
名称Name 基因IDGene ID AA 信号肽Signal peptide PCLD 铜结合位点Copper-binding sites N-糖基化位点N-glycosylation sites ALR GAS DoUCL1 Dendrobium_GLEAN_10023378 148 — 1 H-C-H-M + NC + DoUCL2 Dendrobium_GLEAN_10044449 256 + 1 H-C-H-M + + — DoUCL3 Dendrobium_GLEAN_10077806 128 + 1 H-C-H-M + — — DoUCL4 Dendrobium_GLEAN_10077807 129 + 1 H-C-H-M + — — DoUCL5 Dendrobium_GLEAN_10129350 128 + 1 H-C-H-M + — — DoPLCL1 Dendrobium_GLEAN_10014563 170 + 1 H-C-H-M — + + DoPLCL2 Dendrobium_GLEAN_10014564 170 + 1 H-C-H-M — + + DoPLCL3 Dendrobium_GLEAN_10021763 220 + 1 H-C-H-M — + + DoPLCL4 Dendrobium_GLEAN_10070311 191 + 1 H-C-H-M — + + DoPLCL5 Dendrobium_GLEAN_10077794 128 + 1 H-C-H-M — + — DoPLCL6 Dendrobium_GLEAN_10092500 191 + 1 H-C-H-M — + + DoSCL1 Dendrobium_GLEAN_10027914 207 + 1 H-C-H-Q + + + DoSCL2 Dendrobium_GLEAN_10030280 175 + 1 H-C-H-Q + + + DoSCL3 Dendrobium_GLEAN_10066345 169 + 1 H-C-H-Q + + — DoSCL4 Dendrobium_GLEAN_10066346 169 + 1 H-C-H-Q + + — DoSCL5 Dendrobium_GLEAN_10066347 170 + 1 H-C-H-Q + + — DoSCL6 Dendrobium_GLEAN_10088776 200 — 1 H-C-H-Q + NC + DoSCL7 Dendrobium_GLEAN_10114547 189 + 1 H-C-H-Q + + + DoSCL8 Dendrobium_GLEAN_10136317 199 — 1 H-C-H-Q — NC — DoENODL1 Dendrobium_GLEAN_10008487 133 — 1 — — NC + DoENODL2 Dendrobium_GLEAN_10009350 175 + 1 — — + + DoENODL3 Dendrobium_GLEAN_10011247 187 + 1 — + + + DoENODL4 Dendrobium_GLEAN_10015510 193 + 1 — + + + DoENODL5 Dendrobium_GLEAN_10017023 179 + 1 — + + — DoENODL6 Dendrobium_GLEAN_10028538 204 + 1 — + + — DoENODL7 Dendrobium_GLEAN_10028670 158 + 1 — + + + DoENODL8 Dendrobium_GLEAN_10035898 220 — 1 — + NC + DoENODL9 Dendrobium_GLEAN_10037071 224 + 1 — + + + DoENODL10 Dendrobium_GLEAN_10041716 187 + 1 — + + + DoENODL11 Dendrobium_GLEAN_10045771 358 + 1 — + + + DoENODL12 Dendrobium_GLEAN_10047036 139 — 1 — + NC — DoENODL13 Dendrobium_GLEAN_10059911 220 — 1 — — NC + DoENODL14 Dendrobium_GLEAN_10061798 174 + 1 — + NC + DoENODL15 Dendrobium_GLEAN_10064039 254 + 1 — + + — DoENODL16 Dendrobium_GLEAN_10077812 131 + 1 — — — — DoENODL17 Dendrobium_GLEAN_10077815 187 + 1 — + + — DoENODL18 Dendrobium_GLEAN_10080274 179 + 1 — + + — DoENODL19 Dendrobium_GLEAN_10129392 173 + 1 — + + + AA:氨基酸序列的长度;Length of amino acid sequences; PCLD:(plastocyanin-like domain)PCLD结构域数量;ALR: (Arabinogalactan protein-like region)阿拉伯半乳聚糖样域;GAS:(C-terminal glycosylphosphatidylinositol signal)糖基磷脂酰肌醇锚定信号;+预测到; -未预测到;+ Predicted; -not predicted;NC:由于蛋白序列前端未预测到信号肽而未做阿拉伯半乳聚糖样域的预测; There was no prediction of arabinogalactan protein-like region. -
为进一步明确DoPCs家族成员序列结构特征,本文采用生物信息学分析方法预测了该家族成员蛋白骨架的N-端SP、ALR和GAS以及N-糖基化位点等。预测分析结果显示:有31条DoPCs骨架含有N-端SP,负责引导该链合成的蛋白质进入分泌通路[3]。采用big-PI Plant Predictor和PSORT在线软件预测结果显示:22条DoPCs含有GAS位点,而糖基磷脂酰肌醇锚定蛋白是一种膜蛋白,它可通过糖基磷脂酰肌醇插入内质网膜和质膜,主要负责携带蛋白定位至质膜上,与膜上的脂筏连接[22-23]。此外,有27条DoPCs含N-糖基化位点,可连接糖链[6]。
已知非邻接重复出现的Pro可以被羟基化作为阿拉伯半乳聚糖糖基化位点,而连续重复出现的Pro被羟基化为阿拉伯糖糖基化位点[5, 16]。当Ala、Ser、Thr、Gly和Pro在核心蛋白中为主要氨基酸组分,且Pro以基序为:[Ala/Ser/Thr/Gly]-Pro-X(0, 10)-[Ala/Ser/Thr/Gly]-Pro的非邻接重复方式出现,即为假定AG糖模块[17-19]。有研究发现,当Pro以基序为Ser-Pro2-4连续重复出现时,为假定的伸展蛋白糖模块;然而,Ser-Pro2-4基序中的Pro不完全羟基化时,可形成非邻接型的Pro模块[20]。在拟南芥的相关研究中把高度相似的非邻接型Ala/Thr/Gly-Pro基序和Ser-Pro基序均作为AG糖模块的代表。Mashifuchi等将[Ala/Ser/Thr/Gly]-Pro3-4基序也作为了假定AG糖模块[6]。本文结合前人的研究成果,将[Ala/Ser/Thr/Gly]-Pro-X(0, 10)-[Ala/Ser/Thr/Gly]-Pro和[Ala/Ser/Thr/Gly]-Pro3-4基序作为假定AG糖模块的预测规则。由于N-端SP作为分泌信号,是阿拉伯半乳聚糖蛋白(Arabinogalactan protein,AGP)的先决条件,因此,本文只对含有SP的31条DoPCs是否存在AG糖模块进行预测。经统计,共有26条DoPCs存在AG糖模块,预测为嵌合的AGPs,包括13条DoENODLs、1条DoUCLs、6条DoPLCLs和6条DoSCLs,其中,10条DoPCs预测包含伸展蛋白糖模块,有8条DoPCs既可以是伸展蛋白又可以是AGPs(表 2)。
表 2 DoPCs家族成员阿拉伯半乳糖基模块预测
Table 2. Putative glycomodules in the putative DoPCs
Name 糖基模块Number of putative glycomodules [A/S/T]-P ([A/S/T]-P-X(0, 10)-[A/S/T]-P)X [A/S/T]-P-P [A/S/T]-P-P-P DoUCL1 - - - - DoUCL2 1 1 0 0 DoUCL3 0 0 0 0 DoUCL4 0 0 0 0 DoUCL5 0 0 0 0 DoPLCL1 0 2 0 0 DoPLCL2 0 2 0 0 DoPLCL3 2 8 0 1 DoPLCL4 1 3 1 0 DoPLCL5 2 0 0 0 DoPLCL6 2 2 2 0 DoSCL1 3 2 0 0 DoSCL2 0 2 0 1 DoSCL3 1 0 0 0 DoSCL4 1 0 0 0 DoSCL5 1 0 0 0 DoSCL6 - - - - DoSCL7 2 5 0 0 DoSCL8 - - - - DoENODL1 - - - - DoENODL2 0 0 1 0 DoENODL3 1 4 0 0 DoENODL4 0 3 0 0 DoENODL5 1 0 0 0 DoENODL6 0 5 0 1 DoENODL7 0 0 0 1 DoENODL8 - - - - DoENODL9 0 4 0 0 DoENODL10 1 4 0 0 DoENODL11 0 22 1 2 DoENODL12 - - - - DoENODL13 - - - - DoENODL14 - - - - DoENODL15 1 4 0 0 DoENODL16 0 0 0 0 DoENODL17 1 0 0 0 DoENODL18 1 0 0 0 DoENODL19 0 2 0 0 根据DoPCs和AtPCs蛋白骨架中是否含有N-端SP、PCLD结构域、ALR和C-端GAS这几个部分,将其分为七类(图 2)[5]。类群Ⅰ的DoPCs成员有16个,其蛋白骨架包含所有以上4个结构域和基序;类群Ⅱ和Ⅲ在结构上与类群Ⅰ存在一定的相似,但分别缺少GAS和ALR,其中,归为类群Ⅱ的DoPCs和有10个,归为类群Ⅲ的DoPc有0个;类群Ⅳ同时缺失ALR和C-端GAS两个结构域,有4个成员被归为该类群;归为类群Ⅴ的6个成员和归为类群Ⅵ的2个成员都不存在N-端SP和ALR结构,但前者具有C-端GAS,而后者只具有PCLD这一个结构域;类群Ⅶ同时缺失ALR和C-端GAS,但存在2个PCLD,归为该类群的DoPc成员有0个。由于类群Ⅰ和Ⅱ中存在N-端SP以及足够参与糖基化反应的AG糖模块,可认为这两类中的基因家族成员是嵌合型的AGPs。类群Ⅲ和Ⅶ不含有铁皮石斛PC家族成员,在拟南芥、水稻和杨树中也有类似的现象,这有可能是由于基因在进化过程中失去功能,发生消亡导致[15]。
-
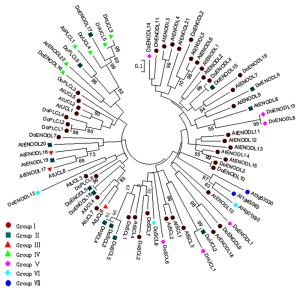
利用已知的38条AtPCs蛋白序列和预测得到的38条DoPCs蛋白序列全长,来构建系统发育树以分析其进化关系。系统发育树结果(图 3)表明:包含SP、PCLD和GAS的类群Ⅲ、包含SP和PCLD的类群Ⅳ及包含SP和2个PCLD的类群Ⅶ的成员较集中,聚集在一起,但每个亚家族的多数成员较为分散地聚集在一起。DoPLCL1/2、DoUCL1/2、DoUCL3/5、DoSCL3/4/5、DoENODL5/18、DoENODL3/10、DoENODL8/13、DoENODL4/15等8对旁系同源蛋白序列相似性较高,并且处于较近的分支上,表明它们之间很可能有功能冗余的现象。
-
为确定DoPC基因在铁皮石斛种子与菌根菌共生萌发过程中的表达水平,本文利用铁皮石斛体外非共生萌发种子(Do)和共生萌发种子(DoTcg)转录组数据进行DoPC基因的表达分析。结果(图 4)表明:16个DoPCs基因在与菌根真菌共生萌发的铁皮石斛种子中检测到表达,其中,DoUCL2、4、DoSCL6、7和DoENOD 3、7、14、17基因表达量较高,而DoUCL5和DoENODL1、17基因在非共生萌发的种子中未检测到表达。对比DoPCs基因在共生萌发和非共生萌发种子中的表达水平,除去3个在非共生萌发种子中未表达的基因,差异表达基因中大部分基因在共生萌发的种子中明显上调表达,只有DoENODL6基因下调表达。这些差异表达基因可能参与铁皮石斛种子与美孢胶膜菌共生萌发生物学调控。
-
通过生物信息学分析,本文对铁皮石斛PC基因家族进行全基因组范围的预测分析,获得38个DoPCs基因。研究报道的拟南芥AtPC、水稻OsPC、大白菜BrPC分别有38、62和84个PC基因[5-7],铁皮石斛基因组DoPC基因数量远少于水稻和大白菜的PC基因,与拟南芥的PC数量相同。对比分析NCBI(http://www.ncbi.nlm.nih.gov)上的基因组数据,拟南芥、水稻和大白菜分别有33 583、30 534和41 147个蛋白编码基因,铁皮石斛基因组35 567个蛋白编码基因,推测该基因家族的数量可能不与基因组蛋白编码基因的数量成正比。研究发现,在拟南芥、水稻和大白菜中,片段复制和串联复制可能是导致PC基因家族扩张的主要原因[5, 7, 15]。
分析该家族的蛋白骨架结构特点发现,2个Cys在DoPCs家族中高度保守,在AtPCs、OsPCs和BrPCs中也寻找到类似的现象,研究发现,2个Cys形成的二硫键有助于维持PCLD结构的稳定,且Cys的存在对于植物PCs家族基因调控功能的发挥有重要意义[5-8]。在铁皮石斛、拟南芥、大白菜等植物中均发现,ENODLs亚家族在对应的铜结合位点上His、Cys、His、Met/Gln已知配体的保守氨基酸残基序列被其它氨基酸残基取代的现象,这些保守氨基酸残基配体很可能与结合铜的功能相关[6-7]。38个DoPCs分为4个亚家族,又根据PC蛋白骨架结构组成特征划分的7个类群,在铁皮石斛PC蛋白序列中未预测到具有类群Ⅲ和Ⅶ特征成员,在拟南芥、水稻和杨树中也有类似的现象,这有可能是由于基因在进化过程中失去功能,发生消亡导致[15]。系统发育分析显示,与拟南芥、水稻、大白菜PCs的相关研究结果一样,各亚家族成员分散聚集在不同分支中,说明所有亚家族成员在单子叶植物和双子叶植物分化发生之前有着共同的祖先,即使在铁皮石斛和拟南芥逐渐分化之后,各亚家族之间也仍在发展进化[5-7]。
阿拉伯半乳聚糖蛋白、伸展蛋白和富脯氨酸蛋白被认为是一大类富羟脯氨酸糖蛋白(Hydroxyproline-rich glycoproteins,HRGPs)。研究表明,HRGPs参与了细胞生长和发育的多个方面,从细胞壁结构的形成到细胞增殖、细胞间识别等[21-22]。铁皮石斛中PC样阿拉伯半乳聚糖蛋白(PLAs)的研究发现,近一半的DoPCs也是嵌合的AGPs,与拟南芥、水稻相比,铁皮石斛中的PC-AGPs数量相对较少[5, 7]。在经典型AGPs的核心蛋白骨架中,氨基酸残基Ala、Ser、Thr、Gly和Hyp是主要构成成分,该区域与C-端的疏水性GAS区域一起,可以使AGP定位到质膜上的脂筏结构域[19, 23]。根据是否包含SP、ALR、GAS和PCLD结构域,共有16个DoPC-PLAs(即类群Ⅰ成员)被预测为经典型的AGPs,而另外10条DoPC-AGPs预测为非经典型蛋白。同时,一些DoPC-AGPs预测到含有伸展蛋白糖模块,作为阿拉伯低聚糖的连接位点。AGPs可能参与细胞分裂和模式形成、幼苗生长、花粉管伸长、植物与微生物互作、植物有性生殖过程等生命活动的调控,其功能的发挥也取决于这些糖蛋白的糖基化情况[24-26]。
PC家族ENODL亚家族基因首先是在豆科植物的根瘤中发现[27],后来发现一些ENODLs基因在植物中受到丛枝菌根真菌侵染的激发而活跃表达[26]。本研究发现,一些DoUCL、DoSCL和DoENODL基因在与菌根菌共生萌发的铁皮石斛种子中高表达,且DoUCL5和DoENODL1、17基因在非共生萌发的铁皮石斛种子中未检测到表达。与之前研究报道的在兰科植物与菌根真菌的共生体中也发现PC家族基因差异化高表达的现象相类似[28],说明不仅ENODL基因在植物与微生物的共生关系可能发挥重要作用,UCL和SCL两个亚家族基因在兰科植物与菌根菌共生关系建立过程中也发挥重要作用。对比DoPCs基因在共生萌发和非共生萌发种子中的表达水平,发现差异表达的PC家族基因中大部分基因在共生萌发的种子中明显上调表达,只有DoENODL6基因明显下调表达,这些在共生萌发种子中差异表达明显的DoPCs可能以不同方式参与铁皮石斛种子与菌根菌共生萌发的生物学调控。
-
本研究在铁皮石斛全基因组中共预测到38个DoPCs,可分属4个亚家族,2个Cys残基在DoPCs家族中高度保守;19条DoPCs蛋白序列含有4个完整的保守铜结合配体(His、Cys、His、Gln/Met),有32条DoPCs骨架中含有N-端信号肽,22条DoPCs含有糖基磷脂酰肌醇锚定位点,有27条DoPCs含N-糖基化位点;16个DoPCs基因在与美孢胶膜菌共生萌发的种子中检测到表达,其中,DoUCL2, 4和DoENODL14基因表达量最高,7个基因在共生萌发的种子中明显上调表达,只有DoENODL6基因明显下调表达。本研究在全基因组范围内分析铁皮石斛DoPC家族蛋白结构特点,可以为探究兰科植物与微生物共生的相互作用及进一步解析兰科菌根形成的分子机制提供数据基础。
铁皮石斛phytocyanin基因家族全基因组分析
Genome-wide Analysis of Phytocyanin Gene Family in Dendrobium officinale
-
摘要:
目的 探讨Phytocyanins(PCs)在铁皮石斛发育过程和逆境胁迫环境下的潜在功能。 方法 利用拟南芥和水稻phytocyanin基因家族蛋白序列(AtPCs和OsPCs), 采用本地化软件BLASTP对铁皮石斛全基因组数据库进行搜索, 并采用SMART和pfam数据库验证质体蓝素样结构域, 获得铁皮石斛phytocyanin基因家族编码序列(DoPCs); 利用Signal P4.1、Big-PI Plant Predictor、NetNGlyc1.0 Server等在线软件对DoPC家族序列的结构特点进行分析; 利用软件ClustalW和MEGA对DoPC基因进行氨基酸序列的比对和系统进化分析; 利用转录组数据绘制heatmap图对DoPCs基因在与美孢胶膜菌共生萌发的种子中的表达情况进行分析。 结果 显示:铁皮石斛全基因组中共预测到包含PCLD结构域的38个DoPCs, 可分属4个亚家族, 2个Cys残基在DoPCs家族中高度保守; 19条DoPCs蛋白序列含有4个完整的保守铜结合配体(His、Cys、His、Gln/Met), 有32条DoPCs骨架中含有N-端信号肽, 22条DoPCs含有糖基磷脂酰肌醇锚定位点, 有27条DoPCs含N-糖基化位点; 16个DoPCs基因在与美孢胶膜菌共生萌发的种子中检测到表达, 其中, DoUCL2, 4和DoENODL14基因表达量最高, 7个基因在共生萌发的种子中明显上调表达, 只有DoENODL6基因明显下调表达。 结论 在全基因组范围内分析铁皮石斛DoPC家族蛋白结构特点, 可以为探究兰科植物与微生物共生的相互作用及进一步研究兰科菌根形成的分子机制提供数据基础。 -
关键词:
- 铁皮石斛
- / phytocyanin基因家族
- / 全基因组分析
- / 共生萌发种子
Abstract:Objective To study the sequence characteristics of Dendrobium officinale Method Local BLASTP was used to search the phytocyanin gene of D. officinale(DoPCs) in the database of Dendrobium whole genome, and then the conserved Plastocyanin-like domain and structural characteristics of DoPCs were predicted. The sequence alignment and phylogenetic analysis of DoPCs were conducted by using Clustal W and MEGA. Finally, 38 DoPCs were expected to have conserved PCLD domains and two Cys residues, belonging to four subfamilies. Result Among these DoPC proteins, 19 DoPCs contained four copper ligands (His, Cys, His, and Gln/Met), 32 DoPCs were predicted having N-terminal signal peptides, 22 DoPCs had putative C-terminal glycosylphosphatidylinositol-anchor signals, and 27 DoPCs had putative arabinogalactan glycomodules. The results of DoPCs expression analysis in symbiotic with Tulasnella calospora and asymbiotic germinated seeds from D. officinale by using RNA-seq data showed that 16 DoPCs were expressed in symbiotic germinated seeds, and DoUCL2, 4 and DoENODL14 had the highest expression with different levels. Seven DoPC genes were obviously up-regulated and only one was down-regulated in symbiotic germinated seeds comparing with that of asymbiotic. Conclusion The study of the protein structure characteristics of DoPCs family in Dendrobium officinale during the whole genome, would help in researching the interaction between Orchidaceae and microorganism, furthermore would serve as data foundation, in researching the molecular mechanism of Orchid mycorrhiza growth. -
表 1 DoPC家族成员及其结构信息
Table 1. The putative DoPCs named information
名称Name 基因IDGene ID AA 信号肽Signal peptide PCLD 铜结合位点Copper-binding sites N-糖基化位点N-glycosylation sites ALR GAS DoUCL1 Dendrobium_GLEAN_10023378 148 — 1 H-C-H-M + NC + DoUCL2 Dendrobium_GLEAN_10044449 256 + 1 H-C-H-M + + — DoUCL3 Dendrobium_GLEAN_10077806 128 + 1 H-C-H-M + — — DoUCL4 Dendrobium_GLEAN_10077807 129 + 1 H-C-H-M + — — DoUCL5 Dendrobium_GLEAN_10129350 128 + 1 H-C-H-M + — — DoPLCL1 Dendrobium_GLEAN_10014563 170 + 1 H-C-H-M — + + DoPLCL2 Dendrobium_GLEAN_10014564 170 + 1 H-C-H-M — + + DoPLCL3 Dendrobium_GLEAN_10021763 220 + 1 H-C-H-M — + + DoPLCL4 Dendrobium_GLEAN_10070311 191 + 1 H-C-H-M — + + DoPLCL5 Dendrobium_GLEAN_10077794 128 + 1 H-C-H-M — + — DoPLCL6 Dendrobium_GLEAN_10092500 191 + 1 H-C-H-M — + + DoSCL1 Dendrobium_GLEAN_10027914 207 + 1 H-C-H-Q + + + DoSCL2 Dendrobium_GLEAN_10030280 175 + 1 H-C-H-Q + + + DoSCL3 Dendrobium_GLEAN_10066345 169 + 1 H-C-H-Q + + — DoSCL4 Dendrobium_GLEAN_10066346 169 + 1 H-C-H-Q + + — DoSCL5 Dendrobium_GLEAN_10066347 170 + 1 H-C-H-Q + + — DoSCL6 Dendrobium_GLEAN_10088776 200 — 1 H-C-H-Q + NC + DoSCL7 Dendrobium_GLEAN_10114547 189 + 1 H-C-H-Q + + + DoSCL8 Dendrobium_GLEAN_10136317 199 — 1 H-C-H-Q — NC — DoENODL1 Dendrobium_GLEAN_10008487 133 — 1 — — NC + DoENODL2 Dendrobium_GLEAN_10009350 175 + 1 — — + + DoENODL3 Dendrobium_GLEAN_10011247 187 + 1 — + + + DoENODL4 Dendrobium_GLEAN_10015510 193 + 1 — + + + DoENODL5 Dendrobium_GLEAN_10017023 179 + 1 — + + — DoENODL6 Dendrobium_GLEAN_10028538 204 + 1 — + + — DoENODL7 Dendrobium_GLEAN_10028670 158 + 1 — + + + DoENODL8 Dendrobium_GLEAN_10035898 220 — 1 — + NC + DoENODL9 Dendrobium_GLEAN_10037071 224 + 1 — + + + DoENODL10 Dendrobium_GLEAN_10041716 187 + 1 — + + + DoENODL11 Dendrobium_GLEAN_10045771 358 + 1 — + + + DoENODL12 Dendrobium_GLEAN_10047036 139 — 1 — + NC — DoENODL13 Dendrobium_GLEAN_10059911 220 — 1 — — NC + DoENODL14 Dendrobium_GLEAN_10061798 174 + 1 — + NC + DoENODL15 Dendrobium_GLEAN_10064039 254 + 1 — + + — DoENODL16 Dendrobium_GLEAN_10077812 131 + 1 — — — — DoENODL17 Dendrobium_GLEAN_10077815 187 + 1 — + + — DoENODL18 Dendrobium_GLEAN_10080274 179 + 1 — + + — DoENODL19 Dendrobium_GLEAN_10129392 173 + 1 — + + + AA:氨基酸序列的长度;Length of amino acid sequences; PCLD:(plastocyanin-like domain)PCLD结构域数量;ALR: (Arabinogalactan protein-like region)阿拉伯半乳聚糖样域;GAS:(C-terminal glycosylphosphatidylinositol signal)糖基磷脂酰肌醇锚定信号;+预测到; -未预测到;+ Predicted; -not predicted;NC:由于蛋白序列前端未预测到信号肽而未做阿拉伯半乳聚糖样域的预测; There was no prediction of arabinogalactan protein-like region. 表 2 DoPCs家族成员阿拉伯半乳糖基模块预测
Table 2. Putative glycomodules in the putative DoPCs
Name 糖基模块Number of putative glycomodules [A/S/T]-P ([A/S/T]-P-X(0, 10)-[A/S/T]-P)X [A/S/T]-P-P [A/S/T]-P-P-P DoUCL1 - - - - DoUCL2 1 1 0 0 DoUCL3 0 0 0 0 DoUCL4 0 0 0 0 DoUCL5 0 0 0 0 DoPLCL1 0 2 0 0 DoPLCL2 0 2 0 0 DoPLCL3 2 8 0 1 DoPLCL4 1 3 1 0 DoPLCL5 2 0 0 0 DoPLCL6 2 2 2 0 DoSCL1 3 2 0 0 DoSCL2 0 2 0 1 DoSCL3 1 0 0 0 DoSCL4 1 0 0 0 DoSCL5 1 0 0 0 DoSCL6 - - - - DoSCL7 2 5 0 0 DoSCL8 - - - - DoENODL1 - - - - DoENODL2 0 0 1 0 DoENODL3 1 4 0 0 DoENODL4 0 3 0 0 DoENODL5 1 0 0 0 DoENODL6 0 5 0 1 DoENODL7 0 0 0 1 DoENODL8 - - - - DoENODL9 0 4 0 0 DoENODL10 1 4 0 0 DoENODL11 0 22 1 2 DoENODL12 - - - - DoENODL13 - - - - DoENODL14 - - - - DoENODL15 1 4 0 0 DoENODL16 0 0 0 0 DoENODL17 1 0 0 0 DoENODL18 1 0 0 0 DoENODL19 0 2 0 0 -
[1] Ruan X, Luo F, Li D, et al. Cotton BCP genes encoding putative blue copper-binding proteins are functionally expressed in fiber development and involved in response to high-salinity and heavy metal stresses[J]. Physiologia Plantarum, 2011, 141(1):71-83. doi: 10.1111/ppl.2011.141.issue-1 [2] Hart P J, Eisenberg D, Nersissian A M, et al. A missing link in cupredoxins-crystal structure of cucumber stellacyanin at 1.6 resolution[J]. Protein Science, 1996, 5(11):2175-2183. doi: 10.1002/pro.v5:11 [3] Nersissian A M, Immoos C, Hill M G, et al. Uclacyanins, stellacyanins, and plantacyanins are distinct subfamilies of phytocyanins Plant-specific mononuclear blue copper proteins[J]. Protein Science, 1998, 7(9):1915-1929. doi: 10.1002/pro.v7:9 [4] Van Driessche G, Van Beeumen J, Dennison C, et al. Heterogeneity of the covalent structure of the blue copper protein umecyanin from horse-radish roots[J]. Protein Science, 1995, 4(2):209-227. [5] Ma H, Zhao H, Liu Z, et al. The phytocyanin gene family in rice (Oryza sativa L.):genome-wide identification, classification and transcriptional analysis[J]. PLoS ONE, 2011, 10:e25184. [6] Mashiguchi K, Asami T, Suzuki Y. Genome-wide identification, structure and expression studies, and mutant collection of 22 early nodulin-like protein genes in Arabidopsis[J]. Bioscience Biotechnology and Biochemistry, 2009, 73(11):2452-2459. doi: 10.1271/bbb.90407 [7] Li J, Gao G, Zhang T, et al. The putative phytocyanin genes in Chinese cabbage (Brassica rapa L.):genome-wide identification, classification and expression analysis[J]. Molecular Genetics and Genomics. 2013, 288(1-2):1-20. doi: 10.1007/s00438-012-0726-4 [8] Dong J, Kim S T, Lord E M. Plantacyanin plays a role in reproduction in Arabidopsis[J]. Plant Physiology, 2005, 138(2):778-789. doi: 10.1104/pp.105.063388 [9] Diab A A, Teulat-Merah B A, This D, et al. Identification of drought-inducible genes and differentially expressed sequence tags in barley[J]. Theoretical and Applied Genetics, 2004, 109(7):1417-1425. doi: 10.1007/s00122-004-1755-0 [10] Ezaki B, Sasaki K, Matsumoto H, et al. Functions of two genes in aluminium (Al) stress resistance:repression of oxidative damage by the AtBCB gene and promotion of efflux of Al ions by the NtGDI1 gene[J]. Journal of Experimental Botany, 2005, 56:2661-2671. doi: 10.1093/jxb/eri259 [11] Wu H, Shen Y, Hu Y, et al. A phytocyanin-related early nodulin-like gene, BcBCP1, cloned from Boea crassifolia enhances osmotic tolerance in transgenic tobacco[J]. Journal of Plant Physiology, 2011, 168(9):935-943. doi: 10.1016/j.jplph.2010.09.019 [12] Zhao M, Zhang G, Zhang D, Hsiao Y, Guo S. ESTs Analysis Reveals Putative Genes Involved in Symbiotic Seed Germination in Dendrobium officinale[J]. PLoS ONE, 2013, 8:e72705. doi: 10.1371/journal.pone.0072705 [13] Zhao X, Yang J, Liu S, et al. The colonization patterns of different fungi on roots of Cymbidium hybridum plantlets and their respective inoculation effects on growth and nutrient uptake of orchid plantlet[J]. World Journal of Microbiology and Biotechnology, 2014, 30(7):1993-2003. doi: 10.1007/s11274-014-1623-2 [14] Deng W, Wang Y, Liu Z, et al. HemI:A Toolkit for Illustrating Heatmaps[J]. PLoS One, 2014, 9:e111988. doi: 10.1371/journal.pone.0111988 [15] Cao J, Li X, Lv Y, et al. Comparative analysis of the phytocyanin gene family in 10 plant species:a focus on Zea mays[J]. Frontiers in Plant Science, 2015, 6:515. [16] Tan L, Leykam J F, Kieliszewski M J. Glycosylation motifs that direct arabinogalactan addition to arabinogalactan-proteins[J]. Plant Physiology, 2003, 132(3):1362-1369. doi: 10.1104/pp.103.021766 [17] Schultz C J, Rumsewicz M P, Johnson K L, et al. Using Genomic resources to guide research directions. The arabinogalactan protein gene family as a test case[J]. Plant Physiology, 2002, 129(4):1448-1463. doi: 10.1104/pp.003459 [18] Schultz C J, Ferguson K L, Lahnstein J, et al. Post-translational modifications of arabinogalactan-peptides of Arabidopsis thaliana[J]. The Journal of Biological Chemistry, 2004, 279(44):45503-45511. doi: 10.1074/jbc.M407594200 [19] Estevez J M, Kieliszewski M J, Khitrov N, et al. Characterization of synthetic hydroxyproline-rich proteoglycans with arabinogalactan protein and extensin motifs in Arabidopsis[J]. Plant Physiology, 2006, 142(2):458-470. doi: 10.1104/pp.106.084244 [20] Shpak E, Barbar E, Leykam J F, et al. Contiguous hydroxyproline residues direct hydroxyproline arabinosylation in Nicotiana tabacum[J]. The Journal of Biological Chemistry, 2001, 276(14):11272-11278. doi: 10.1074/jbc.M011323200 [21] Majewska-Sawka A, Nothnagel E. The multiple roles of arabinogalactan proteins in plant development[J]. Plant Physiology, 2000, 122(1):3-9. doi: 10.1104/pp.122.1.3 [22] Showalter A M. Arabinogalactan-proteins:structure, expression and function[J]. Cellular and Molecular Life Sciences, 2001, 58(10):1399-1417. doi: 10.1007/PL00000784 [23] Borner G H H, Lilley K S, Stevens T J, et al. Identification of glycosylphosphatidylinositol-anchored proteins in Arabidopsis. A proteomic and genomic analysis[J]. Plant Physiology, 2003, 132(2):568-577. doi: 10.1104/pp.103.021170 [24] De Vries S C. A relationship between seed development, arabinogalactan-proteins (AGPs) and the AGP mediated promotion of somatic embryogenesis[J]. Physiologia Plantarum, 2002, 114(4):637-644. doi: 10.1034/j.1399-3054.2002.1140418.x [25] Tan L, Showalter A M, Egelund J, et al. Arabinogalactan-proteins and the research challenges for these enigmatic plant cell surface proteoglycans[J]. Frontiers In Plant Science, 2012, 3(140):1-10. [26] Harrison M J. Development of the arbuscular mycorrhizal symbiosis[J]. Current Opinion in Plant Biology, 1998, 1(4):360-365. doi: 10.1016/1369-5266(88)80060-8 [27] Nap J P, Bisseling T. Noduling function and nodulin gene regulation in root nodule development[M]//Gresshoff P M. The Molecular Biology of Symbiotic Nitrogen Fixation. Boca Raton: CRC Press Inc, 1980: 181-229. [28] Perotto S, Rodda M, Benetti A, et al. Gene expression in mycorrhizal orchid protocorms suggests a friendly plant-fungus relationship[J]. Planta, 2014, 239(6):1337-1349. doi: 10.1007/s00425-014-2062-x -
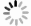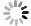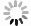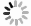



 下载:
下载: