-
合理的林分密度能够显著促进林木的径向生长[1-2],从而提高木材产量,解析种植密度促进林木径向生长的内在机制意义重大。林木的径向生长(即次生木质部生成)来源于维管形成层的活动。维管形成层细胞向外分化产生次生韧皮部,向内分化产生次生木质部,其中,次生木质部细胞不断积累,使植株径向增粗[3]。由维管形成层分化产生成熟的次生木质部细胞需要经历木质部母细胞分化和分裂、木质部细胞增大、次生壁加厚、细胞程序化凋亡和心材形成多个阶段[4],各个阶段在分子水平上均受到一大批基因的调控。因此,研究木质部在不同种植密度下的基因表达模式,挖掘和鉴定响应种植密度的木质部生成关键基因并解析其作用机制是研究种植密度影响林木径向生长内在机制的重要内容。
以往以模式植物拟南芥(Arabidopsis thaliana (Linn.) Heynh.)和杨属(Populus)为研究对象,分离鉴定了一些参与木质部生成各个阶段的关键调控基因,在木质部生成方面取得了重要进展。诸多研究表明,调控因子如受体激酶和转录因子等在木质部生成过程中发挥重要的调控作用,如PTL(PETAL LOSS,花瓣缺失)转录因子作为抑制子参与调控拟南芥根系发育过程中的形成层细胞分裂和木质部细胞分化活动[5];类受体激酶PXY和其家族成员PXL1、PXL2参与调控拟南芥维管组织的细胞分裂活动[6-7];类受体激酶FEI和SHOU参与次生壁物质合成,fei突变体纤维素合成减少,而SHOU能够恢复fei突变体表型[8-9]。次生壁合成也需要一些转录因子参与,Chen等[10]在毛果杨(Populus trichocarpa Torr. & Gray)中绘制了一个4层级的细胞壁合成转录调控网络,其中包含17个转录因子和27个细胞壁成分合成基因,它们协同调控木质部生成;最近在杨属、松属(Pinus)等树种中又相继证实了多个NAC和MYB家族成员参与木材细胞壁物质合成过程[11-13]。由此可见,次生木质部生成的分子机制十分复杂,仍然有许多未知亟待解析。
桉属(Eucalyptus)与杨属为世界公认的速生人工林树种,但二者木材的生物量和质量存在明显差异。这2个树种在木质部生成调控机制上可能存在差异,但目前关于桉属植物木质部生成相关重要基因的解析及其调控机制的研究相对较少[14-15],同时,关于种植密度影响径向生长的分子基础研究尚未见报道。因此,本研究以不同种植密度下尾巨桉(Eucalyptus urophylla × Eucalyptus grandis W.Hill ex Maiden)的主茎为试验材料,联合PacBio Iso-Seq和RNA-Seq转录组技术,通过对尾巨桉木质部差异表达转录本(DETs)的表达模式展开分析,挖掘和鉴定桉树中响应种植密度的木质部生成关键基因,以期为科学阐明种植密度影响林木径向生长的分子机制奠定理论基础。
HTML
-
试验地位于广东省湛江市南方国家级林木种苗示范基地内(111°38′ E,21°30′ N)。试验材料为长势均一、主茎通直、无病虫害的尾巨桉DH32-29无性系组织培育苗。2019年7月14日,将幼苗移植至高0.5 m直径0.3 m的种植袋中,并按照10株·m−2高种植密度(株行距0.3 m × 0.3 m)和5株·m−2低种植密度(株行距0.6 m × 0.6 m)移放至长 × 宽 × 高为5.0 m × 4.5 m × 0.5 m的样地内,每个处理设置3个重复,共计6个样地。在幼苗生长过程中,水分和营养充分供应且保持一致。苗木种植后定期测定生长性状,于直径及其增长率表现出显著差异的时期进行取样(2019年10月15日)(表1)。每个样地选择1株平均木,取距离地面10 cm处主茎的木质部、韧皮部,共计12个样本。液氮冷冻后,带回试验室置于−80℃的超低温冰箱保存备用。样品命名为H-X1~3、L-X1~3、H-P1~3、L-P1~3,其中,H为高种植密度,L为低种植密度,X表示木质部,P表示韧皮部,数字1~3代表生物学重复。
处理
Treatment平均直径/mm
Average diameter直径生长量/mm
Diameter increment平均株高/cm
Average height株高生长量/cm
Height increment低种植密度 Low planting density 6.84 ± 0.19 a 2.36 ± 0.28 a 65.72 ± 2.64 a 18.80 ± 2.51 a 高种植密度 High planting density 5.41 ± 0.20 b 1.41 ± 0.15 b 58.53 ± 2.21 a 13.97 ± 2.92 a 注:不同小写字母表示不同种植密度间的生长性状具有显著性差异(P=0.01)。
Note: The different letters indicate growth traits with significant differences in different planting densities(P=0.01).Table 1. Effect of planting density on growth traits for seedlings of E. urophylla × E. grandis
-
使用植物RNA提取试剂盒DP130508(天根生物公司)分别提取12个样本的RNA,等量混合后使用SMARTer™ PCR cDNA Synthesis Kit试剂盒合成mRNA的全长cDNA,通过PacBio平台进行全长转录组测序(百迈客公司);使用CD-HIT软件[16]去除转录本中的冗余序列;使用BLAST软件(version 2.2.26)[17]将得到非冗余转录本序列和NR、eggNOG、Pfam、GO、Swissprot、KOG、COG、KEGG数据库比对进行转录本功能注释;使用iTAK软件[18]进行调控因子预测。
对12个样本的RNA分别进行cDNA合成和文库构建,委托百迈客公司进行Illumina二代测序。使用RSEM[19]软件,通过比对到三代转录本上的位置信息,对转录本的表达水平进行定量,采用FPKM(Fragments Per Kb of transcript per Million fragments mapped)作为衡量指标。
-
使用DESeq2[20]对样本H-X1、H-X2、H-X3和L-X1、L-X2、L-X3进行分析,获得2个种植密度间DETs。筛选标准为差异倍数(Fold Change) ≥ 2且错误发现率(FDR)< 0.01;使用百迈客云平台热图绘制工具绘制DETs 热图,每个表达量的数值均取以10为底数的对数(Log10)[21];使用STRING数据库[22]对DETs 构建蛋白互作网络。
-
以转录组试验中2个种植密度下尾巨桉幼苗的木质部RNA为模板,使用TSK302S RT6 cDNA Synthesis Kit Ver 2(擎科生物公司)合成cDNA模板,从443个DETs中随机选取9个DETs,进行qRT-PCR验证转录组测序结果的准确性;选择9个DETs进行组织表达分析,材料为尾巨桉幼苗的叶片、韧皮部、第2节间木质部(木质部2)和第7节间木质部(木质部7)4个部位,RNA的提取采用植物RNA提取试剂盒V1.5(百菲特生物公司)。利用Primer 5.0设计qRT-PCR引物,以arp4(Actin-related protein 4)编码基因作为内参,引物见表2。使用2 × T5 Fast qPCR Mix(SYBR Green I)荧光定量试剂盒(擎科生物公司)进行荧光定量检测。每个样品设置4个技术重复,采用2−ΔΔCt计算相对表达量。使用SPSS 25.0处理数据,使用Excel 2016软件绘图,以P < 0.01表示差异显著。
序号
Number转录本编号
Transcripts ID上游引物
Forward primer下游引物
Reverse primer1 MIX-PB_transcript_46917 GGAGATGAAGAATGGCGTGC CAGCGGAAGAACTCGCATAC 2 MIX-PB_transcript_39150 GGGACTAAAATTATCGGATTACGGT GCATAGTCTGTTGATGTTGGTAGCA 3 MIX-PB_transcript_72449 GTTCCTGGTCTCCGACAGTT AGGCAAGATACAGCACCCTG 4 MIX-PB_transcript_15209 CTCTGAAATCGGTGACTTCCTC AAGCATCTTTGCCACCCAAC 5 MIX-PB_transcript_17152 TCGTGGATGCGTTGCTTACT GGGCTCTCGGGTTCCTAATG 6 MIX-PB_transcript_61822 CAAACGAGTGGGAGCAAGGA CGGATCTCAATGCCATGCTG 7 MIX-PB_transcript_39624 TCGAGCACGCATTTTCTGTTT GGGTCGCAGGTGGTTCTATT 8 MIX-PB_transcript_6405 GTTGGATTTGGGTCGTGTCG GAGACTAGACACACCGGACAG 9 MIX-PB_transcript_6089 CCTTCGCTAAGAACAATCCACG AGCTTGCGGGTACCATGTCATA 10 MIX-PB_transcript_67597 CGACCAGATTCCAGGAGAGC GCCAGATCGAGACACCTTAAACT 11 MIX-PB_transcript_18228 GCTTTCTCCCCACAAGAGGA TCGTGTTTGGCGAGTGTCTA 12 MIX-PB_transcript_51497 GCCACCAAAGGACCGGAATA ACTTCTCTTGGCACTCACCG 13 MIX-PB_transcript_15184 GGCGGGAAAAGAAAAGCACG GCGAAATCAGAGACGCTCAC 14 MIX-PB_transcript_70069 CCTGATGATACGGGCTTCGT ATGATCCGTCACTCTTGCTGT 15 MIX-PB_transcript_73425 AGCTGGAATGTTGTAGTCGCT CTCAAGGTCCACATCTGCCAA 16 MIX-PB_transcript_12050 GTTTCCTGGCCGATAACCCA AGGAGGAGGGGTCAACACTT 17 MIX-PB_transcript_52911 GGTCATGGAGTGTGTCAAGC TTCCCACGCCATTGTAAGAGA 18 MIX-PB_transcript_30100 TGCTGATGCTTGACAGACAG AAGAAGCACCCACAATCCAC 19 arp4 GGAATTGTCGCGGATTGGGA GCCTTTTCTCGTTGCTGCTG Table 2. Primers for qRT-PCR
1.1. 试验材料
1.2. 转录组测序及结果分析
1.3. DETs筛选及分析
1.4. DETs实时荧光定量PCR验证
-
对高、低2个种植密度下尾巨桉主茎木质部和韧皮部进行Pacbio全长转录组测序,共获得247 536条环形一致序列(CCS),其中,209 654条为全长非嵌合序列;对全长非嵌合序列进行聚类分析,获得81 317条一致性序列;进一步对一致性序列进行校正,最终获得高质量转录本(准确度>99%)79 965条,低质量转录本1 281条,高质量转录本达到98%以上;经CD-HIT去冗余后,获得非冗余转录本序列45 490条(表3)。
分类
Classification统计
CountCCS数量
Number of CCS247 536 全长非嵌合序列数量
Number of full-length non-chimeric reads209 654 一致性序列数量
Number of consensus isoforms81 317 高质量转录本数量
Number of polished high-quality isoforms79 965 低质量转录本数量
Number of polished low-quality isoforms1 281 高质量转录本所占比例
Percent of polished high-quality isoforms/%98.34 非冗余转录本数
Number of non-redundant transcripts45 490 Table 3. Statistical table of PacBio Iso-Seq data evaluation
对高、低两个种植密度下尾巨桉主茎木质部进行RNA-Seq测序,共获得130 292 914条150 bp双端Reads。6个样本中,测序获得最多的Clean Reads为23 322 440条,最少为19 998 315条;获得Clean Data碱基数最多为6 988 132 460 bp,最少为5 990 895 036 bp;GC含量为50.30%~51.05%,%≥Q30(Clean Data质量值≥30的碱基所占的百分比)为93.88%~94.41%(表4),GC含量和%≥Q30均值分别为50.60%、94.19%。
样品
SamplesClean Reads数量
Number of Clean ReadsClean Data碱基数/bp
Number of Clean DataGC含量/%
GC Content大于或等于Q30的百分比/%
≥ Q30H-X1 21 882 942 6 557 229 728 50.52 94.09 H-X2 22 886 461 6 843 134 076 50.81 94.41 H-X3 21 093 958 6 317 393 546 51.05 94.40 L-X1 19 998 315 5 990 895 036 50.42 94.21 L-X2 21 108 798 6 323 257 814 50.52 94.16 L-X3 23 322 440 6 988 132 460 50.30 93.88 共计/均值
Total/Mean130 292 914 39 020 042 660 50.60 94.19 Table 4. Statistical table of RNA-Seq data evaluation
-
将三代转录组测序获得的45 490个非冗余转录本与NR、eggNOG、Pfam、GO、Swissprot、KOG、COG、KEGG数据库比对并进行功能注释,有44 888条转录本至少比对到一个数据库中,注释率达98.68%(表5),其中,NR数据库注释率达98.47%,eggNOG、Pfam、Swissprot、GO、KOG数据库注释到的转录本数量均超过50%,KEGG数据库注释到的转录本数量最低。
数据库
Annotated databases注释到的转录本数目
Number of transcripts注释率/%
Annotation rateNR 44854 98.47 eggNOG 43433 95.47 Pfam 37496 82.43 GO 36631 80.52 Swissprot 33276 73.15 KOG 27767 61.04 COG 19986 44.93 KEGG 17093 37.58 All 44888 98.68 Table 5. Statistics of BLAST annotation of different databases
-
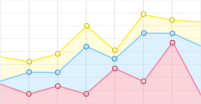
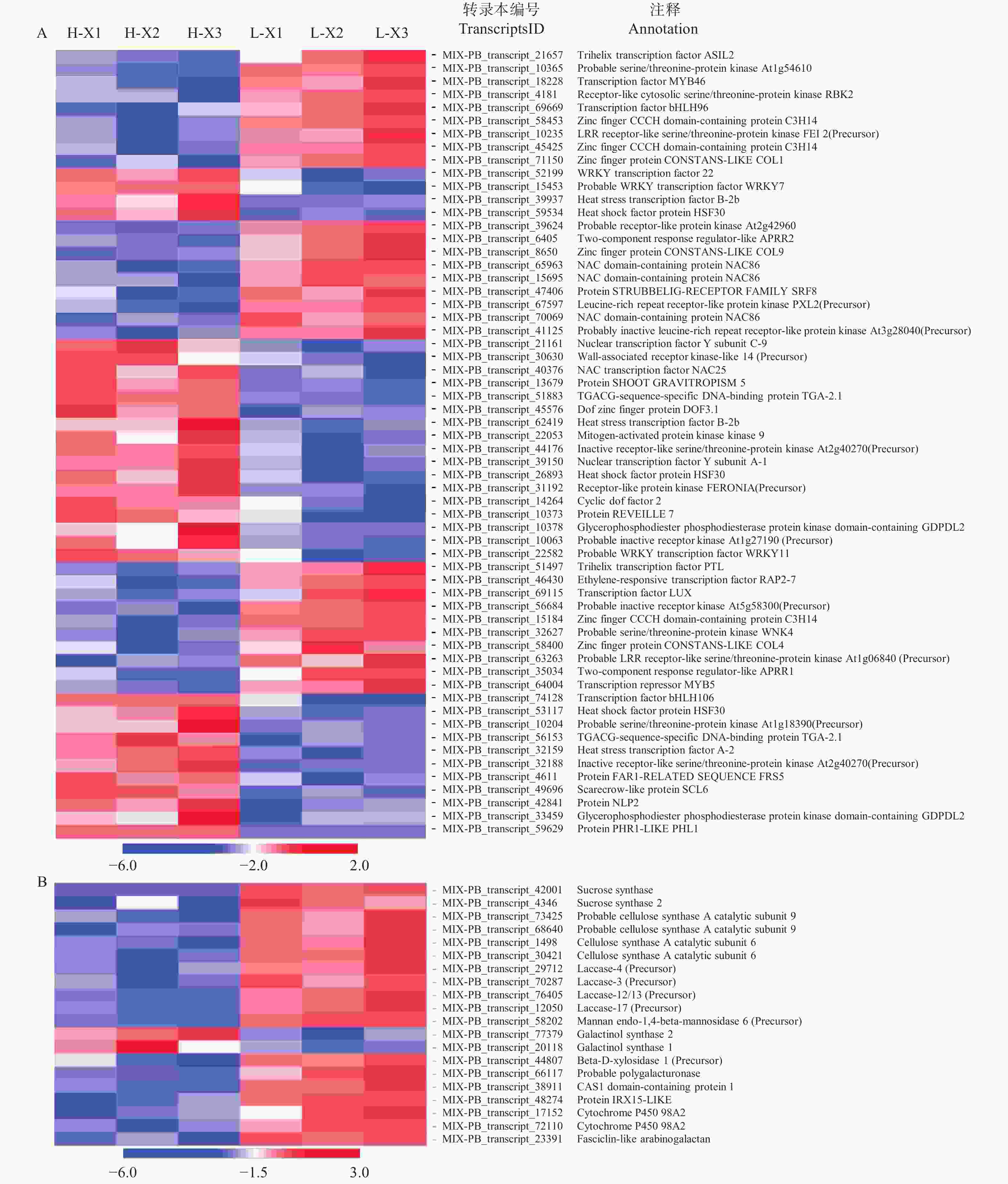
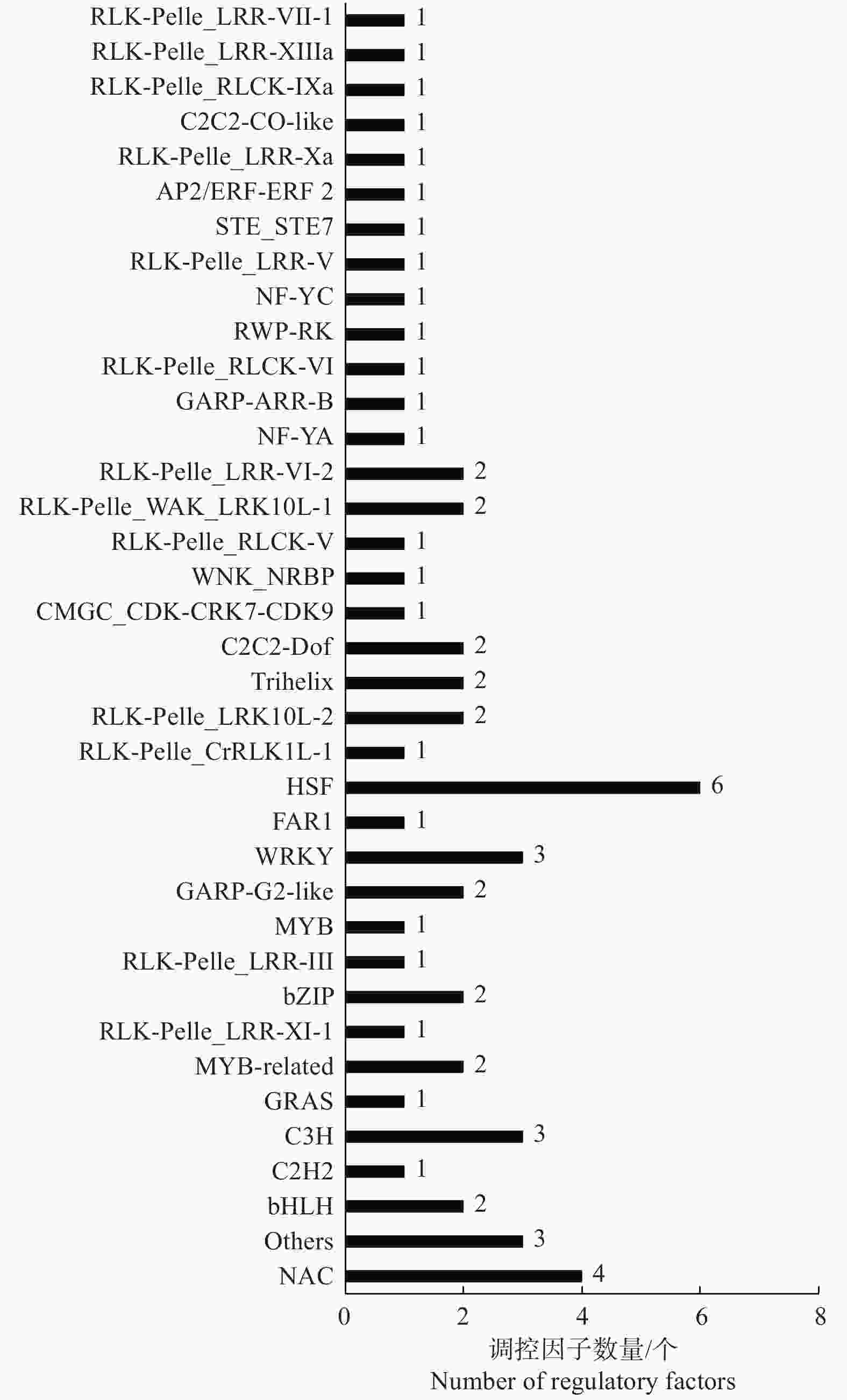
为了充分了解不同种植密度下尾巨桉木质部转录本的表达模式,以H-X为对照组,L-X为试验组,筛选获得差异表达转录本443个,其中,215个DETs在低种植密度条件下上调表达,228个DETs下调表达。注释结果显示,443个DETs中包含了60个假定的调控因子,归为37个家族(图1A、图2),其中,28个差异表达调控因子在低种植密度下尾巨桉的木质部中上调表达,编码21类调控因子。这些调控因子与多种生物学过程有关,GARP-ARR-B、Trihelix和RLK-Pelle_LRR-XI-1参与调控维管形成层活动;C3H、MYB、NAC和RLK-Pelle_LRR-XIIIa参与调控细胞壁成分合成;RLK-Pelle_LRR-III参与次生生长过程;RLK-Pelle_RLCK-V参与细胞周期调控;GARP-G2-like和WNK_NRBP参与节律调控。32个DETs下调表达,所对应调控因子的功能多与外界胁迫有关,如热激转录因子HSF、参与盐胁迫的bHLH和STE_STE7、参与干旱胁迫的RLK-Pelle_WAK_LRK10L-1、参与病虫害应答的RLK-Pelle_LRR-VI-2和RLK-Pelle_LRR-Xa、对脱落酸(ABA)敏感的bHLH和RLK-Pelle_CrRLK1L-1等。除编码调控因子的转录本外,一些转录本的表达模式也发生了显著的变化,在低种植密度植株的木质部中细胞壁物质合成相关蛋白(蔗糖合成酶,纤维素合成酶,漆酶,糖基转移酶,糖基水解酶)和木质素物质合成相关编码基因多为上调表达(图1B)。
-
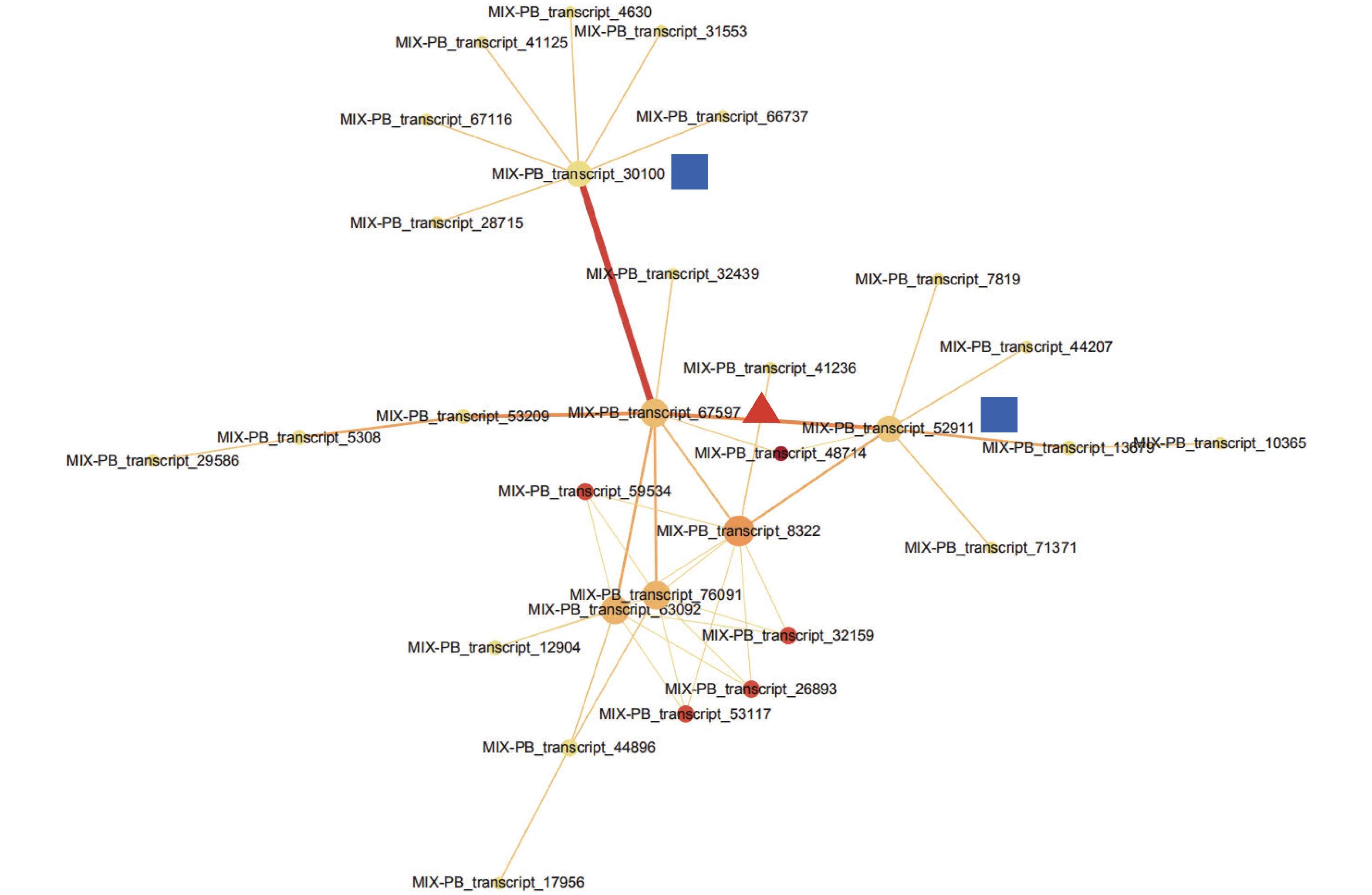
通过STRING数据库对443个DETs进行蛋白互作网络分析,检测到一簇相互作用的基因簇,共包含30个转录本(图3,附表1)。这些基因主要与参与翻译后修饰、蛋白质转换和伴侣蛋白,细胞周期控制、细胞分裂和染色体分割,转录等功能。MIX-PB_transcript_67597作为网络的核心基因,与MIX-PB_transcript_30100和MIX-PB_transcript_52911占据了网络的中心位置(图3)。数据库注释表明:MIX-PB_transcript_67597编码类受体激酶家族RLK-Pelle_LRR-XI-1的PXL2,注释功能为转录。转录本MIX-PB_transcript_52911编码蛋白CUL1参与目标蛋白的泛素化;MIX-PB_transcript_30100编码蛋白T15D22.7参与硫醇氧化酶和硫氧还蛋白家族蛋白的合成,二者在数据库的注释功能均为细胞周期控制、细胞分裂和染色体分割。这3个基因在低种植密度植株的木质部中均上调表达,可能在尾巨桉主茎径向生长中发挥关键作用(附表1)。
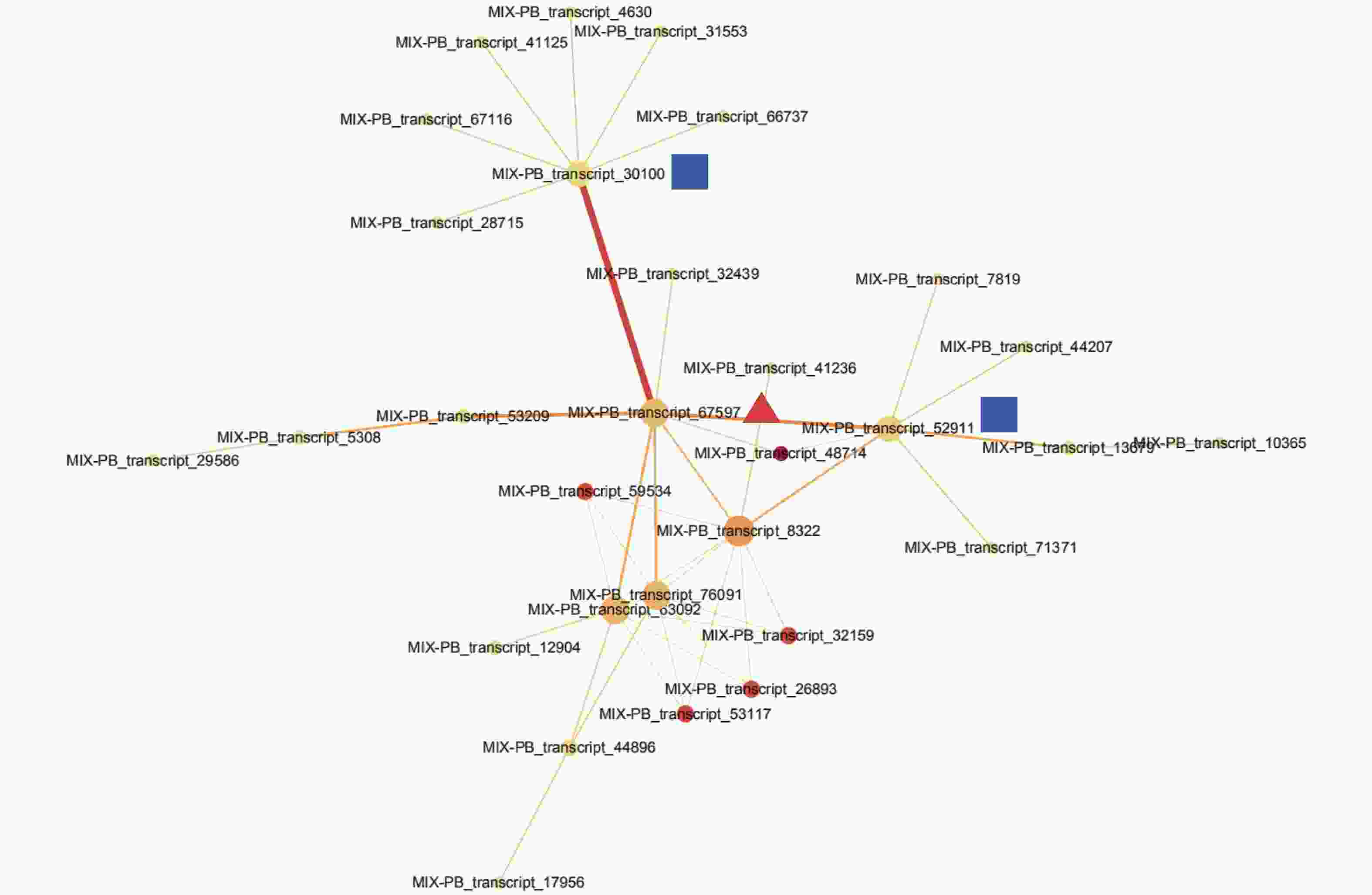
Figure 3. The interaction network of proteins of differentially expressed transcripts in xylem for E. urophylla × E. grandis under different planting densities
转录本 Transcripts ID 转录本的功能 Transcript function 注释Annotation MIX-PB_transcript_67597 转录
Transcription富含亮氨酸的重复类受体蛋白激酶PXL2(前体)
Leucine-rich repeat receptor-like protein kinase PXL2 (Precursor)MIX-PB_transcript_52911 细胞周期控制,细胞分裂,染色体分割
Cell cycle control, cell division, chromosome partitioning类枯灵素蛋白1
cullin-1-like ,CUL1MIX-PB_transcript_30100 细胞周期控制,细胞分裂,染色体分割
Cell cycle control, cell division, chromosome partitioning巯基氧化酶1(前体)
Sulfhydryl oxidase 1 (Precursor), T15D22.7MIX-PB_transcript_31553 -- 推定的抗病蛋白RGA4 Putative disease resistance protein RGA4 MIX-PB_transcript_4630 -- Toll/白细胞介素-1受体-类蛋白
Toll/interleukin-1 receptor-like protein, TIRMIX-PB_transcript_41125 -- 可能的无活性富亮氨酸重复受体类蛋白激酶
Probably inactive leucine-rich repeat receptor-like protein kinase At3g28040MIX-PB_transcript_67116 -- 烟草花叶病毒抗性蛋白N
TMV resistance protein NMIX-PB_transcript_66737 -- 烟草花叶病毒抗性蛋白N
TMV resistance protein NMIX-PB_transcript_28715 -- 烟草花叶病毒抗性蛋白N
TMV resistance protein NMIX-PB_transcript_32439 信号转导机制
Signal transduction mechanisms可能的蛋白磷酸酶2C60
Probable protein phosphatase 2CMIX-PB_transcript_41236 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperones类硫氧还蛋白1-1,叶绿体(前体)
Thioredoxin-like 1-1, chloroplastic (Precursor)MIX-PB_transcript_53209 -- 金属内蛋白酶1(前体)
Metalloendoproteinase 1 (Precursor)MIX-PB_transcript_5308 -- 枯草杆菌蛋白酶样蛋白酶(前体)
Subtilisin-like protease (Precursor)MIX-PB_transcript_29586 脂质运输和代谢Lipid transport and metabolism C型1类尼曼-匹克蛋白
Niemann-Pick C1 protein isoform X1MIX-PB_transcript_48714 一般功能预测General function prediction only 锚定重复序列蛋白
Ankyrin repeat-containing protein, At5g02620MIX-PB_transcript_59534 转录Transcription 热激因子蛋白30
Heat shock factor protein HSF30MIX-PB_transcript_8322 转录Transcription 热激蛋白83
Heat shock protein 83MIX-PB_transcript_76091 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperones热激同源蛋白70
Heat shock cognate 70 kDa protein HSP70MIX-PB_transcript_63092 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperones热激同源蛋白70
Heat shock cognate 70 kDa protein HSP70MIX-PB_transcript_12904 一般功能预测General function prediction only 多元醇转运蛋白5
Polyol transporter 5, PLT5MIX-PB_transcript_32159 转录Transcription 热应激转录因子A-2 T19L18.4
Heat stress transcription factor A-2 T19L18.4MIX-PB_transcript_26893 转录Transcription 热激因子蛋白30
Heat shock factor protein HSF30MIX-PB_transcript_53117 转录Transcription 热激因子蛋白30
Heat shock factor protein HSF30MIX-PB_transcript_44896 氨基酸转运和代谢
Amino acid transport and metabolism类酪氨酸脱羧酶1
tyrosine decarboxylase 1-likeMIX-PB_transcript_17956 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperonesDnaJ同源蛋白(前体)
DnaJ protein homolog (Precursor), DNAJ1MIX-PB_transcript_7819 一般功能预测
General function prediction onlyF框/亮氨酸精氨酸精氨酸重复蛋白3
F-box/LRR-repeat protein 3, FBL3MIX-PB_transcript_44207 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperonesE3泛素蛋白连接酶环1
E3 ubiquitin-protein ligase RING1MIX-PB_transcript_13679 一般功能预测General function prediction only 顶端向地性蛋白5 Protein SHOOT GRAVITROPISM 5 MIX-PB_transcript_10365 细胞周期控制,细胞分裂,染色体分割
Cell cycle control, cell division, chromosome partitioning可能的丝氨酸/苏氨酸蛋白激酶
Probable serine/threonine-protein kinase, At1g54610MIX-PB_transcript_71371 翻译后修饰,蛋白质转换,伴侣蛋白
Posttranslational modification, protein turnover, chaperonesE3泛素蛋白连接酶环1
E3 ubiquitin-protein ligase RING1Table 1. The annotation of 30 differentially expressed transcripts responding to planting density in protein interaction network
-
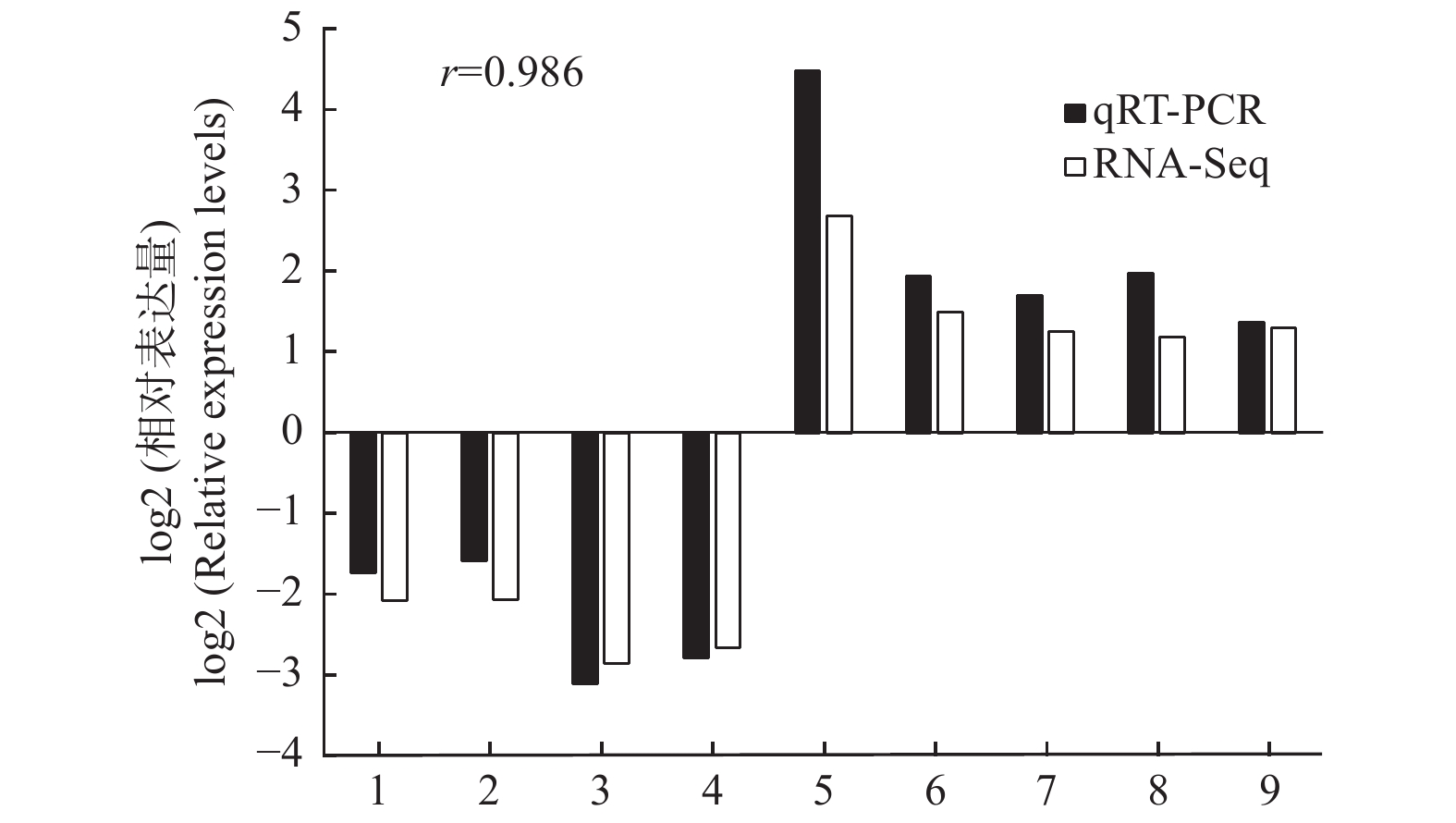
以转录组测序中2个种植密度下的木质部RNA为模板,随机选择9个DETs进行qRT-PCR验证,qRT-PCR结果与转录组测序的表达结果基本一致(图4),证明转录组测序数据准确可靠。
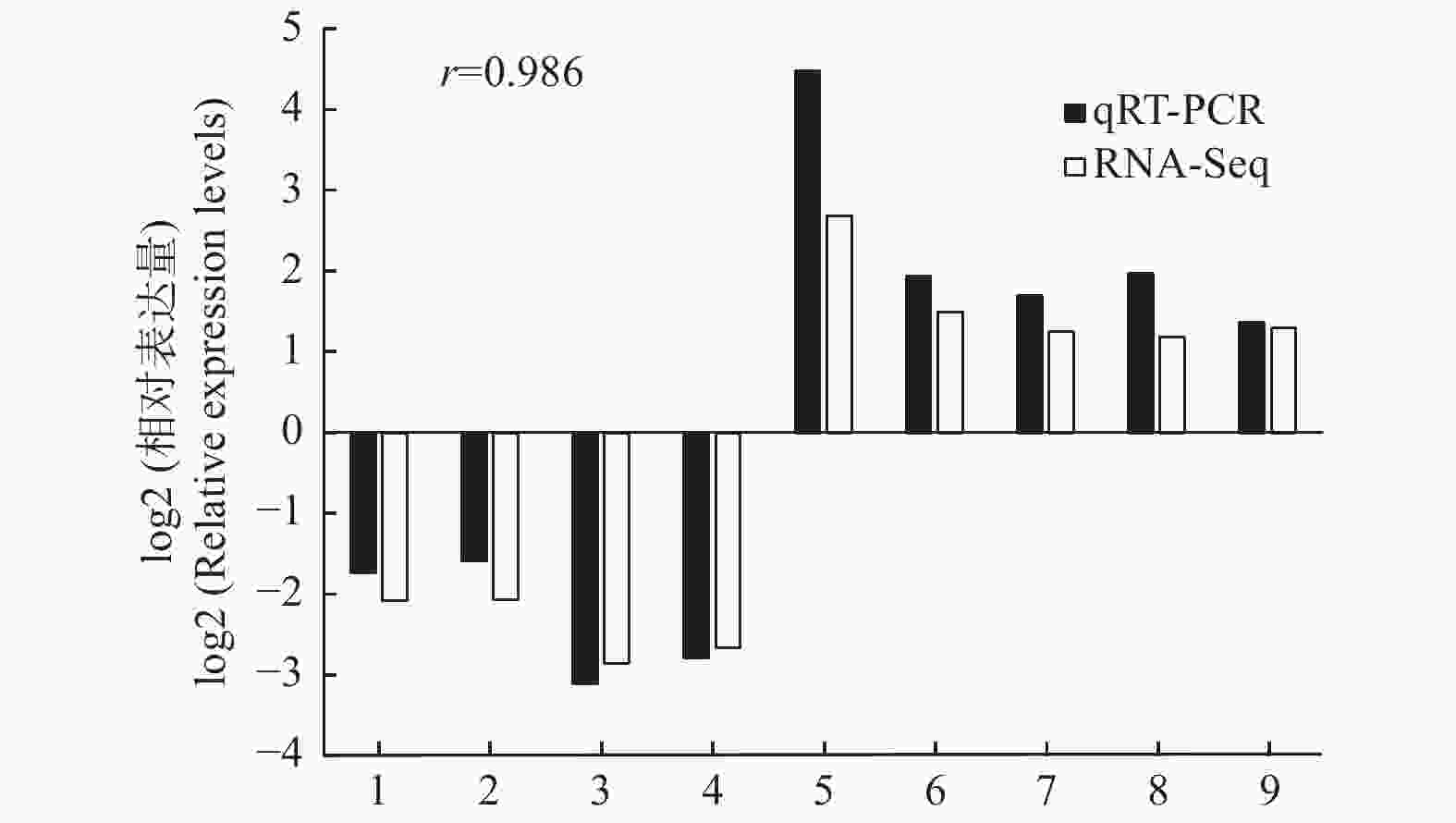
Figure 4. Confirmation of the expression profiles of 9 differentially expressed transcripts by qRT-PCR
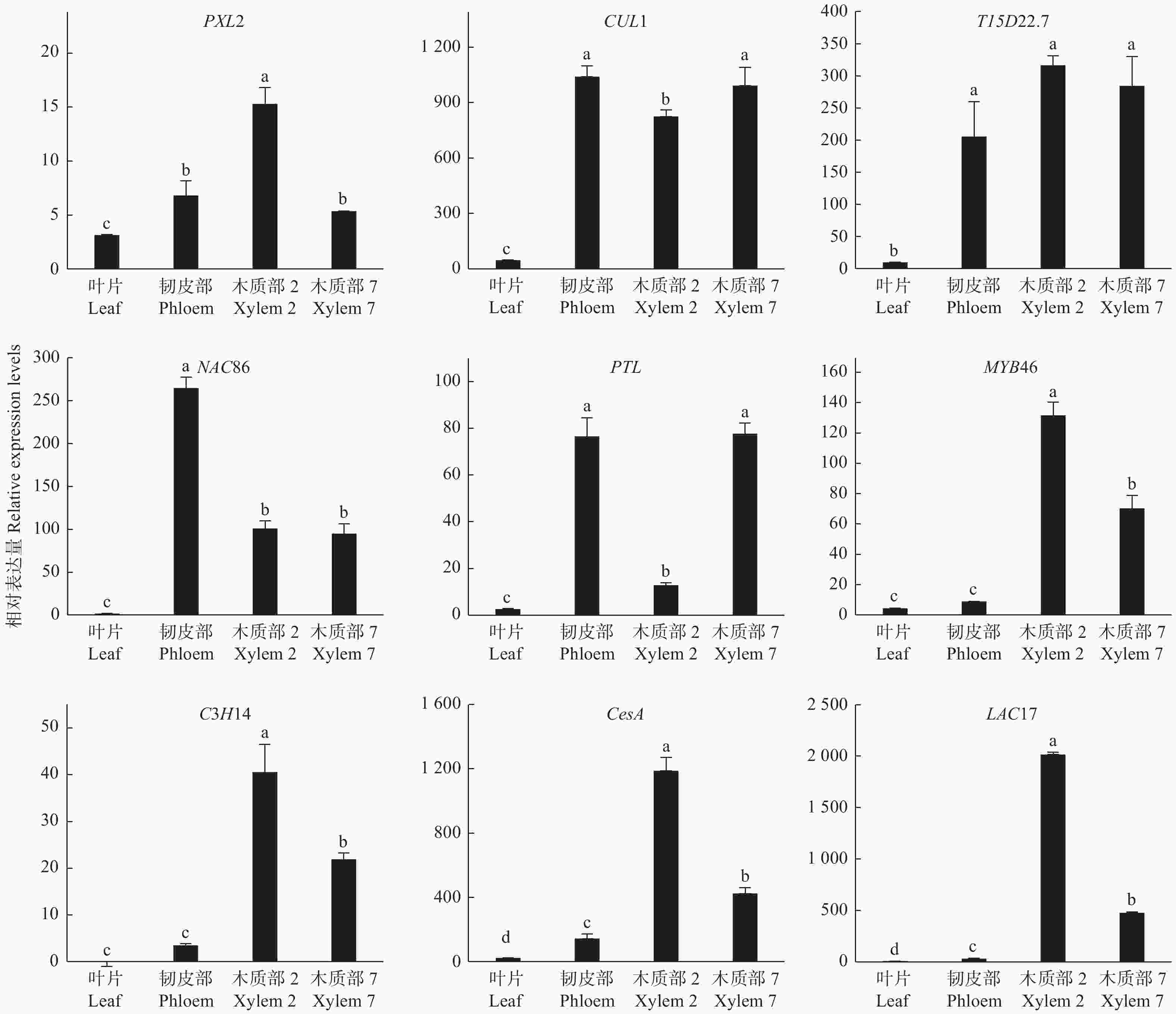
为了验证木质部生成候选基因的功能,通过qRT-PCR对9个木质部生成候选基因进行分析,检测了它们在不同组织的基因表达情况。这9个DETs分别是转录因子基因PTL、MYB46、C3H14、NAC86,细胞壁合成酶基因CesA、LAC17以及PXL2、T15D22.7、CUL1。qRT-PCR结果(图5):这些编码基因在叶片中几乎不表达,在韧皮部、木质部均有不同程度的表达,其中,PXL2在木质部2中的表达量最高,在韧皮部和木质部7中表达水平也较高;CUL1和T15D22.7在韧皮部和木质部中高水平表达;PTL在韧皮部和木质部7中表达量最高,显著高于叶片和木质部2;NAC86在韧皮部中表达水平最高,但在木质部2和木质部7中也有较高表达水平;MYB46、C3H14、CesA和LAC17的表达模式相对一致,在木质部2的表达量最高,木质部7中略低,而韧皮部中表达量极低。这些结果进一步证明了测序结果的可靠性,同时也表明这些基因可能在木质部生成中发挥重要作用。
2.1. 转录组测序统计
2.2. 不同种植密度下尾巨桉主茎转录组分析
2.3. 不同种植密度木质部转录本表达模式分析
2.4. 差异表达转录本蛋白互作网络分析
2.5. qRT-PCR验证
-
为了研究种植密度影响林木径向生长的分子机制,本研究联合PacBio Iso-Seq和RNA-Seq测序技术,在高、低2个种植密度下尾巨桉DH32-29幼苗直径及直径生长量开始出现显著差异时进行转录组测序,获得45 490条在主茎表达的非冗余转录本,共有44 888(98.68%)个转录本得到注释,其中,在NR数据库中有98.47%(44 854/45 490)的转录本能够预测到同源序列,注释率较高,可能是因为巨桉(Eucalyptus grandis Hill)全基因组测序已经完成,剩余602(1.32%)个转录本未得到注释,可能是尾巨桉中的新基因。由于缺乏尾巨桉全基因组序列,本研究结合二代和三代转录组测序结果进行分析有利于数据的分类和定量研究,提高了序列的准确性和可靠性。
木材的生长性状是由多基因决定的高度复杂性状。调控因子在木材形成过程中发挥了重要的调控作用,改变调控因子的表达就可促使多个功能基因共同发挥作用,从而达到性状改良的效果[10, 23]。因此,本研究重点关注木质部中响应种植密度的60个差异表达调控因子。高种植密度植株木质部中优势表达的调控因子多数与胁迫相关,这与欧美杨(Populus × euramericana (Dode) Guinier)中参与胁迫响应的转录因子在高种植密度植株叶片中上调表达相一致[24];而低种植密度中优势表达的调控因子主要与细胞分裂活动和细胞壁物质合成相关。这可能是因为在高种植密度条件下,植株生长过密,易受多种生物或非生物因子的胁迫,从而引起胁迫相关基因上调表达以响应劣势的生长环境,而在低种植密度条件下有利于植株生长,促进了木质部生成相关基因的表达,因而使其径向生长显著大于高种植密度。
为了更好的挖掘和鉴定响应种植密度的木质部生成关键基因,本研究进一步对低种植密度植株木质部中优势表达的调控因子进行分析,发现了一些重要的调控因子编码基因PXL2、MYB46、C3H14、NAC86和PTL,它们在低种植密度植株木质部中均上调表达(图1A)。蛋白互作网络和组织表达分析结果表明,类受体激酶PXL2与2个关键节点基因CUL1和T15D22.7互作并具有一致的表达模式(图3),这与拟南芥中的研究相吻合,PXL2与PXL1、PXY共同协作调控维管组织发育过程中细胞分裂活动[6];此外,杨属植物中PXL2的同源基因PXY通过调控维管组织的细胞分裂活动参与木质部生成[25],由此表明,PXL2、CUL1和T15D22.7可能共同协作调控尾巨桉主茎维管组织的细胞分裂活动,这可能是造成不同种植密度下植株径向生长差异的重要原因之一。
MYB和锌指蛋白C3H是植物中普遍存在的转录因子,越来越多的研究表明,这些转录因子在植物次生壁的合成中发挥重要作用[26-31]。在拟南芥中,MYB46及其杨属和桉属植物中的同源基因PtrMYB21和EgMYB2是次生壁合成调控网络的主控开关成员,能够直接调控其它转录因子和次生壁合成相关基因的表达[26-30];AtC3H14及其杨属植物中同源基因PdC3H17/18是MYB46的直接靶基因,可正向调控次生壁的合成及木质部生成相关MYBs基因的表达[31]。本研究表明,MYB46、C3H14、CesA和参与木质素合成的漆酶基因LAC17等主要在木质部中表达,且均在直径生长量显著增大的植株上调表达,推测MYB46、C3H14可能参与调控LACs和CesAs的表达,并且MYB46、C3H14可能通过参与尾巨桉木质部生成过程中次生壁合成的调控对种植密度做出响应。这些结果表明木质部生成过程中转录调控的复杂性,进一步揭示这些调控因子在木质部生成中的作用具有重要意义。
值得关注的是,在拟南芥根系发育过程中,NAC86直接参与韧皮部内筛管分子成熟过程的调控[32],PTL作为抑制子参与调控木质部细胞分化活动[5],尚未见二者参与木本植物木质部生成调控的报道。在本研究中,NAC86、PTL在低种植密度植株的木质部中上调表达(图1A),在韧皮部和木质部中均有表达(图5),这表明拟南芥同源基因NAC86和PTL可能同时参与木本植物中木质部和韧皮部的生成,PTL可能在木本植物的维管组织发育过程中发挥正向调控作用,暗示这2个基因可能具有与草本植物中不同的功能,值得进一步研究。
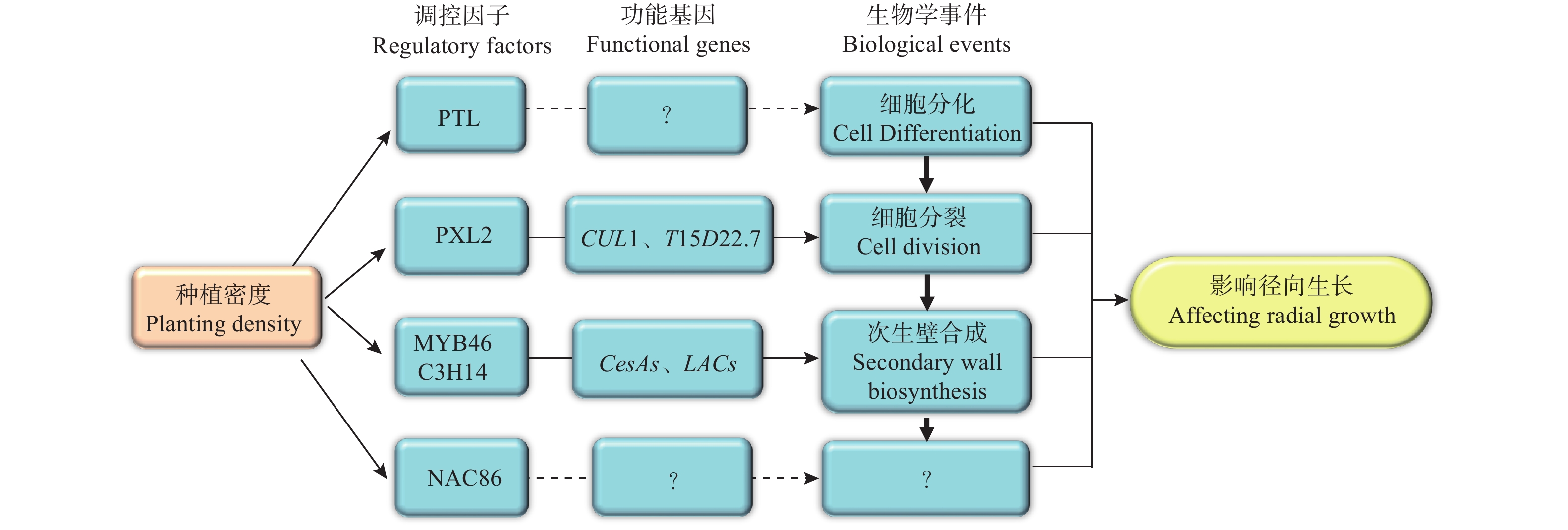
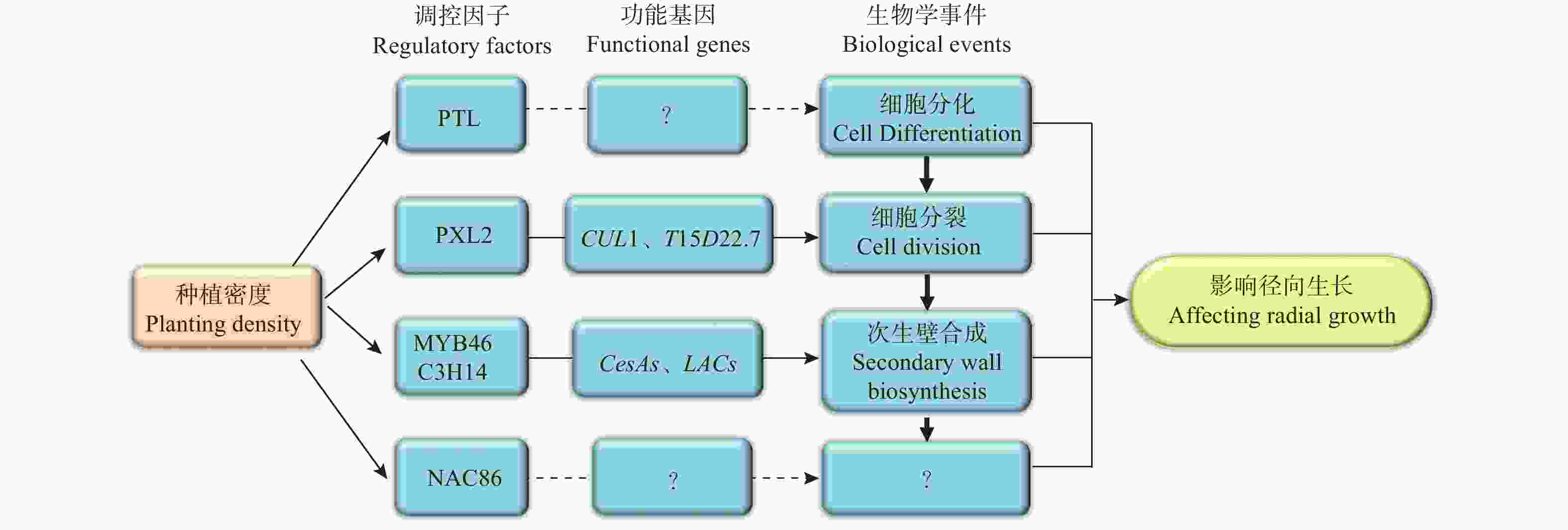
基于以上研究结果,笔者提出了关于种植密度影响尾巨桉径向生长的分子调控网络模型(图6)。该模型中,种植密度通过影响调控维管组织细胞分裂的PXL2、调控次生壁合成的MYB46和C3H14、以及调控木质部细胞分化的PTL及其下游功能基因的表达模式进而促进了植株的径向生长。然而,该模型中各部分成员尤其是PTL、NAC86的具体功能和作用机制仍有待深入研究。为此,接下来将进一步开展基因功能研究,明确模型中关键成员如PTL、NAC86和PXL2在木材形成中的功能,并开展调控机制研究,明确其上下游调控基因,同时将研究结果推广到更多研究群体中,验证研究结论普遍性,以使研究结果得以为生产实践应用。
-
本研究从全基因组转录水平和基因表达两方面,鉴定了参与种植密度响应的显著表达的基因,获得了参与桉树径向生长的候选调控因子,提出了种植密度影响尾巨桉径向生长的分子调控网络模型,并提出NAC86和PTL在木本植物径向生长与草本植物生长中的功能具有差异,该研究对于揭示桉树人工林密度调控的分子机制和木质部生成关键基因的研究具有重要意义。



 DownLoad:
DownLoad: